Valine Degradation
| Source | ID | Pathway |
|---|---|---|
| PathWhiz | PW002489 |
Pathway Metabolites
| YMDB ID | Name CAS Number | IUPAC Name | Formula Weight | Structure |
|---|---|---|---|---|
| YMDB00110 | NAD 53-84-9 | 1-[(2R,3R,4S,5R)-5-[({[({[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl)oxy](hydroxy)phosphoryl}oxy)methyl]-3,4-dihydroxyoxolan-2-yl]-3-carbamoyl-1lambda5-pyridin-1-ylium | C21H28N7O14P2 664.433 | 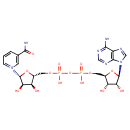 |
| YMDB00143 | NADH 58-68-4 | [({[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl)oxy]({[(2R,3S,4R,5R)-5-(3-carbamoyl-1,4-dihydropyridin-1-yl)-3,4-dihydroxyoxolan-2-yl]methoxy})phosphinic acid | C21H29N7O14P2 665.441 | 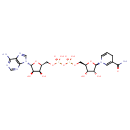 |
| YMDB00152 | L-Valine 72-18-4 | (2S)-2-amino-3-methylbutanoic acid | C5H11NO2 117.1463 | 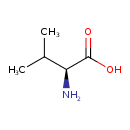 |
| YMDB00153 | Oxoglutaric acid 328-50-7 | 2-oxopentanedioic acid | C5H6O5 146.0981 | 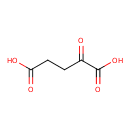 |
| YMDB00271 | L-Glutamic acid 56-86-0 | (2S)-2-aminopentanedioic acid | C5H9NO4 147.1293 | 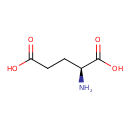 |
| YMDB00365 | Alpha-Ketoisovaleric acid 759-05-7 | 3-methyl-2-oxobutanoic acid | C5H8O3 116.1152 | 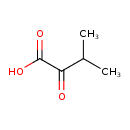 |
| YMDB00482 | isobutyraldehyde 78-84-2 | 2-methylpropanal | C4H8O 72.1057 |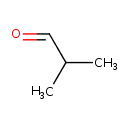  |
| YMDB00573 | isobutanol 78-83-1 | 2-methylpropan-1-ol | C4H10O 74.1216 | 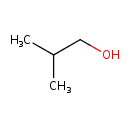 |
| YMDB00862 | hydron 12408-02-5 | hydron | H 1.00794 |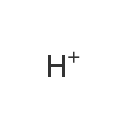  |
| YMDB00912 | Carbon dioxide 124-38-9 | methanedione | CO2 44.0095 | 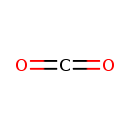 |