| Identification |
|---|
| Name | Asparagine synthetase [glutamine-hydrolyzing] 1 |
|---|
| Synonyms | - Glutamine-dependent asparagine synthetase 1
|
|---|
| Gene Name | ASN1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in asparagine synthase (glutamine-hydrolyzing) activity |
|---|
| Specific Function | ATP + L-aspartate + L-glutamine + H(2)O = AMP + diphosphate + L-asparagine + L-glutamate |
|---|
| Cellular Location | Not Available |
|---|
| SMPDB Pathways | |
|---|
| KEGG Pathways | | Alanine, aspartate and glutamate metabolism | ec00250 |  | | Nitrogen metabolism | ec00910 | 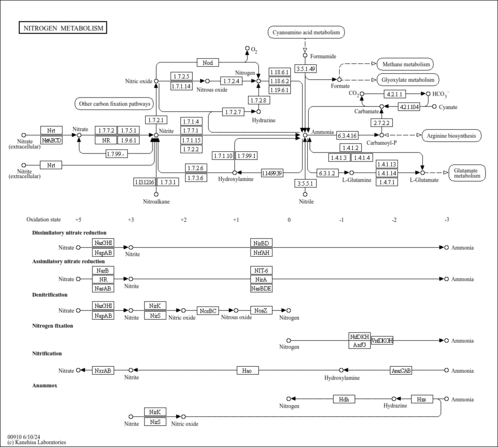 |
|
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00002 | L-Glutamine | Show | | YMDB00097 | Adenosine monophosphate | Show | | YMDB00109 | Adenosine triphosphate | Show | | YMDB00219 | Pyrophosphate | Show | | YMDB00226 | L-Asparagine | Show | | YMDB00271 | L-Glutamic acid | Show | | YMDB00423 | Ammonium | Show | | YMDB00862 | hydron | Show | | YMDB00890 | water | Show | | YMDB00896 | L-Aspartic acid | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| asparagine synthase (glutamine-hydrolyzing) activity | | catalytic activity | | ligase activity | | ligase activity, forming carbon-nitrogen bonds | | carbon-nitrogen ligase activity, with glutamine as amido-N-donor | | Process |
|---|
| cellular amino acid and derivative metabolic process | | cellular amino acid metabolic process | | aspartate family amino acid metabolic process | | asparagine metabolic process | | asparagine biosynthetic process | | metabolic process | | cellular metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 16 |
|---|
| Locus | YPR145W |
|---|
| Gene Sequence | >1719 bp
ATGTGTGGTATTTTCGCCGCTTTCAGGCACGAAGACGTGCATAGATATAAGCCAAAGGCT
CTACAACTATCAAAAAGAATCAGACACCGTGGTCCAGATTGGTCCGGTAATGCTATCAAG
AACTCCACTATATTTGTTCATGAAAGACTAGCCATTGTCGGTGTGGAATCCGGTGCTCAA
CCAATTACTTCTTCAGACGGAGAGTACATGCTATGTGTTAACGGTGAAATCTACAACCAC
ATTCAATTAAGAGAAGAATGCGCAGACTACGAGTTTGGAACACTGAGTGACTGTGAGCCT
ATCATCCCAATGTACTTAAAGCACGATATCGACGCTCCTAAGTACTTGGATGGTATGTTT
GCTTGGACTCTTTACGACGCTAAACAAGATCGTATTGTGGCAGCCAGAGACCCAATCGGT
ATTACGACATTATATATGGGACGCTCTTCCGCTTCTCCAAAGACCGTTTATTTTGCATCC
GAACTAAAATGTTTGACTGACGACTGTGACACTATCACTGCATTCCCACCGGGCCACGTA
TACGATTCTAAGACTGACAAGATCACCCGTTACTTCACACCAGATTGGCTGGACGAAAAA
CGCATTCCTTCCACCCCAATAGATTACATGGCAATTAGACACTCCTTAGAAAAAGCCGTT
AGAAAGAGATTAATGGCCGAAGTCCCATACGGTGTTCTATTGTCGGGTGGTTTGGACTCC
TCTTTAATCGCTTCCATTGCTGCCCGTGAAACTGCAAAGGCCACTAACGATGTCGAACCA
TCAACTTACGATAGTAAGGCAAGACATCTAGCAGGTATCGACGATGACGGTAAGCTACAC
ACTGCTGGTTGGACAAGTCTCCATTCCTTTGCCATCGGTTTACCAAATGCTCCAGATTTG
CAAGCCGCAAGAAAGGTTGCCAAATTCATCGGCTCTATTCATCATGAACACACCTTTACA
TTACAAGAAGGTTTGGATGCTTTGGACGACGTGATCTACCATTTGGAAACTTACGACGTT
ACCACTATCAGAGCTTCCACTCCAATGTTCTTACTATCCAGAAAGATTAAGGCCCAAGGG
GTCAAGATGGTTCTTTCCGGTGAAGGTTCCGATGAAATCTTCGGTGGTTATCTATATTTC
GCACAAGCTCCTTCTGCGGCAGAATTTCACACTGAATCCGTGCAACGTGTCAAGAACTTG
CATTTGGCAGATTGTTTGAGAGCTAACAAGTCTACGATGGCTTGGGGTCTAGAAGCTCGT
GTTCCATTCTTAGACAGAGAATTTTTGCAATTGTGTATGAACATCGATCCAAATGAAAAG
ATGATTAAACCAAAGGAAGGACGTATTGAAAAGTACATTCTAAGGAAGGCATTCGACACC
ACAGGAGAACCAGATGCTAAGCCATATTTACCAGAAGAAATTTTATGGAGACAAAAAGAA
CAATTTTCCGACGGTGTTGGTTACTCCTGGATCGACGGATTAAAAGATACCGCCGAAGCA
GTCATTTCAGATGAAATGTTTGCCAGTCCAAAGGCCGAATGGGGTAGTGACATTCCAACC
ACAAAAGAGGCTTTCTGGTACAGACTAAAATTCGATGCTTTGTTCCCTCAAAAGACCGTG
GCTGACACCGTTATGAGATGGATTCCAAAGGCCGACTGGGGTTGTGCTGAAGATCCTTCT
GGTAGATATGCCCAAATTCATGAAAAACATATCGAATAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 572 |
|---|
| Protein Molecular Weight | 64469.60156 |
|---|
| Protein Theoretical pI | 6.02 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Asparagine synthetase [glutamine-hydrolyzing] 1
MCGIFAAFRHEDVHRYKPKALQLSKRIRHRGPDWSGNAIKNSTIFVHERLAIVGVESGAQ
PITSSDGEYMLCVNGEIYNHIQLREECADYEFGTLSDCEPIIPMYLKHDIDAPKYLDGMF
AWTLYDAKQDRIVAARDPIGITTLYMGRSSASPKTVYFASELKCLTDDCDTITAFPPGHV
YDSKTDKITRYFTPDWLDEKRIPSTPIDYMAIRHSLEKAVRKRLMAEVPYGVLLSGGLDS
SLIASIAARETAKATNDVEPSTYDSKARHLAGIDDDGKLHTAGWTSLHSFAIGLPNAPDL
QAARKVAKFIGSIHHEHTFTLQEGLDALDDVIYHLETYDVTTIRASTPMFLLSRKIKAQG
VKMVLSGEGSDEIFGGYLYFAQAPSAAEFHTESVQRVKNLHLADCLRANKSTMAWGLEAR
VPFLDREFLQLCMNIDPNEKMIKPKEGRIEKYILRKAFDTTGEPDAKPYLPEEILWRQKE
QFSDGVGYSWIDGLKDTAEAVISDEMFASPKAEWGSDIPTTKEAFWYRLKFDALFPQKTV
ADTVMRWIPKADWGCAEDPSGRYAQIHEKHIE |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Dang, V. D., Valens, M., Bolotin-Fukuhara, M., Daignan-Fornier, B. (1996). "Cloning of the ASN1 and ASN2 genes encoding asparagine synthetases in Saccharomyces cerevisiae: differential regulation by the CCAAT-box-binding factor." Mol Microbiol 22:681-692.8951815
- Bussey, H., Storms, R. K., Ahmed, A., Albermann, K., Allen, E., Ansorge, W., Araujo, R., Aparicio, A., Barrell, B., Badcock, K., Benes, V., Botstein, D., Bowman, S., Bruckner, M., Carpenter, J., Cherry, J. M., Chung, E., Churcher, C., Coster, F., Davis, K., Davis, R. W., Dietrich, F. S., Delius, H., DiPaolo, T., Hani, J., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome XVI." Nature 387:103-105.9169875
- Smolka, M. B., Albuquerque, C. P., Chen, S. H., Zhou, H. (2007). "Proteome-wide identification of in vivo targets of DNA damage checkpoint kinases." Proc Natl Acad Sci U S A 104:10364-10369.17563356
- Albuquerque, C. P., Smolka, M. B., Payne, S. H., Bafna, V., Eng, J., Zhou, H. (2008). "A multidimensional chromatography technology for in-depth phosphoproteome analysis." Mol Cell Proteomics 7:1389-1396.18407956
|
|---|

