| Identification |
|---|
| Name | Glutamine-dependent NAD(+) synthetase |
|---|
| Synonyms | - NAD(+) synthase [glutamine-hydrolyzing]
|
|---|
| Gene Name | QNS1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in NAD+ synthase (glutamine-hydrolyzing) activity |
|---|
| Specific Function | ATP + deamido-NAD(+) + L-glutamine + H(2)O = AMP + diphosphate + NAD(+) + L-glutamate |
|---|
| Cellular Location | Cytoplasmic |
|---|
| SMPDB Pathways | |
|---|
| KEGG Pathways | | Nicotinate and nicotinamide metabolism | ec00760 | 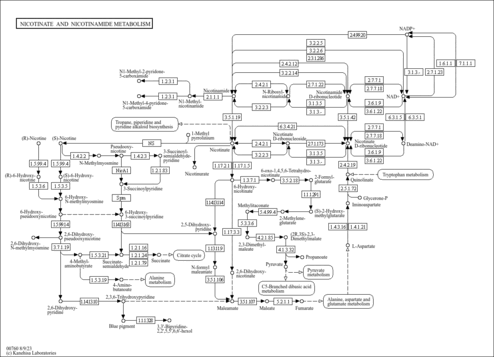 |
|
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00002 | L-Glutamine | Show | | YMDB00097 | Adenosine monophosphate | Show | | YMDB00109 | Adenosine triphosphate | Show | | YMDB00110 | NAD | Show | | YMDB00219 | Pyrophosphate | Show | | YMDB00271 | L-Glutamic acid | Show | | YMDB00306 | deamido-NAD(+) | Show | | YMDB00412 | NAD(+) | Show | | YMDB00423 | Ammonium | Show | | YMDB00862 | hydron | Show | | YMDB00890 | water | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds | | catalytic activity | | NAD+ synthase (glutamine-hydrolyzing) activity | | ligase activity | | ligase activity, forming carbon-nitrogen bonds | | carbon-nitrogen ligase activity, with glutamine as amido-N-donor | | binding | | nucleoside binding | | purine nucleoside binding | | adenyl nucleotide binding | | adenyl ribonucleotide binding | | ATP binding | | hydrolase activity | | Process |
|---|
| metabolic process | | cofactor metabolic process | | coenzyme metabolic process | | coenzyme biosynthetic process | | pyridine nucleotide biosynthetic process | | nitrogen compound metabolic process | | nicotinamide nucleotide biosynthetic process | | NAD biosynthetic process | | cellular metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 8 |
|---|
| Locus | YHR074W |
|---|
| Gene Sequence | >2145 bp
ATGTCACATCTTATCACTTTAGCTACATGCAACTTGAATCAATGGGCCCTAGATTTTGAA
GGTAATAGAGACCGTATCCTACAGTCCATTAAGATTGCCAAAGAGAGGGGTGCCAGGTTA
CGTGTCGGCCCAGAACTGGAAATAACTGGCTACGGATGTTTAGATCATTTTTTAGAAAAT
GACGTTTGCCTTCATTCATGGGAAATGTATGCTCAAATCATTAAGAATAAAGAAACCCAT
GGATTAATACTTGACATTGGTATGCCCGTTCTACACAAGAATGTTCGTTATAATTGTCGT
TTGTTATCCTTGGATGGTGAGATATTGTTCATAAGACCTAAGATTTGGTTAGCTAATGAT
GGTAACTATAGGGAAATGAGATTTTTCACACCTTGGATGAAACCTGGCGTGGTGGAGGAC
TTTATCCTTCCACCTGAGATTCAGAAAGTTACCGGCCAGAGACTTGTGCCATTTGGGGAC
GCTGTGATAAATTCATTGGATACATGCATTGGTACAGAAACTTGTGAAGAATTGTTTACA
CCTCAATCCCCCCACATCGCCATGTCTTTAGATGGTGTGGAAATCATGACAAACTCATCT
GGTTCTCATCATGAACTGCGTAAGTTAAATAAAAGGTTAGACCTAATTTTAAATGCCACT
AAACGTTGTGGTGGTGTTTACTTGTATGCAAATCAAAGAGGTTGTGATGGTGACAGATTA
TATTATGATGGCTGTGCACTAATTGCCATCAATGGTACAATTGTAGCCCAAGGTTCACAA
TTTTCGCTAGATGATGTGGAAGTAGTTACTGCTACTGTGGACCTAGAAGAGGTGAGGAGT
TATCGTGCAGCTGTCATGTCTCGTGGCCTACAAGCCTCCTTGGCAGAAATAAAGTTCAAG
CGTATTGATATTCCTGTAGAATTGGCTTTAATGACCTCCAGATTTGATCCTACAGTGTGT
CCAACAAAAGTCCGCGAGCCTTTCTATCACTCTCCTGAGGAAGAAATTGCACTGGGACCT
GCTTGCTGGATGTGGGATTATTTAAGACGTTGTAACGGAACAGGGTTTTTCCTTCCCTTA
TCTGGGGGCATTGACTCTTGTGCAACTGCAATGATTGTCCACTCTATGTGCCGTTTAGTG
ACCGACGCTGCTCAAAATGGAAATGAGCAAGTTATCAAAGACGTTCGTAAGATAACACGT
AGCGGCGATGATTGGATTCCAGACAGTCCACAGGATCTAGCCTCAAAAATATTTCACTCC
TGTTTCATGGGTACGGAAAATTCATCCAAGGAGACAAGAAACAGAGCAAAGGACCTTTCC
AATGCAATTGGATCTTACCACGTGGATTTAAAGATGGACTCATTGGTATCCAGTGTGGTG
TCCTTATTCGAAGTAGCCACTGGCAAAAAACCAATATACAAAATATTTGGGGGATCTCAA
ATCGAGAACTTGGCTTTACAAAACATCCAGGCGCGTCTAAGAATGGTTCTTTCTTATCTT
TTTGCGCAACTGTTGCCGTGGGTTCGTGGTATCCCAAACTCGGGTGGATTGTTAGTACTT
GGTAGCGCAAATGTTGATGAGTGCTTACGTGGGTATCTAACAAAATATGACTGCTCCTCC
GCAGATATCAACCCTATTGGGGGTATTTCAAAAACTGACTTGAAAAGATTCATTGCCTAC
GCATCAAAACAATATAACATGCCAATCTTGAATGACTTTTTAAACGCTACACCAACTGCA
GAATTAGAACCTATGACTAAAGATTACGTTCAATCGGATGAGATAGATATGGGGATGACG
TATGAAGAATTGGGCGTGTTTGGTTACCTAAGAAAGGTTGAAAAATGTGGTCCTTATTCT
ATGTTCTTAAAACTTCTTCATCAATGGTCCCCAAAGTTAACACCTCGTCAAATATCTGAA
AAGGTGAAAAGATTTTTCTTCTTCTATGCCATCAACAGACACAAGCAAACTGTTTTAACT
CCTAGTTATCATGCTGAACAGTATTCACCAGAAGACAACAGATTTGACTTACGTCCTTTC
TTAATCAACCCAAGATTTCCATGGGCTTCAAGAAAAATTGATGAAGTTGTCGAGCAGTGT
GAAGCACATAAAGGCTCAACGCTTGACATTATGTCTATTGATTAG |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 714 |
|---|
| Protein Molecular Weight | 80684.89844 |
|---|
| Protein Theoretical pI | 6.51 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Glutamine-dependent NAD(+) synthetase
MSHLITLATCNLNQWALDFEGNRDRILQSIKIAKERGARLRVGPELEITGYGCLDHFLEN
DVCLHSWEMYAQIIKNKETHGLILDIGMPVLHKNVRYNCRLLSLDGEILFIRPKIWLAND
GNYREMRFFTPWMKPGVVEDFILPPEIQKVTGQRLVPFGDAVINSLDTCIGTETCEELFT
PQSPHIAMSLDGVEIMTNSSGSHHELRKLNKRLDLILNATKRCGGVYLYANQRGCDGDRL
YYDGCALIAINGTIVAQGSQFSLDDVEVVTATVDLEEVRSYRAAVMSRGLQASLAEIKFK
RIDIPVELALMTSRFDPTVCPTKVREPFYHSPEEEIALGPACWMWDYLRRCNGTGFFLPL
SGGIDSCATAMIVHSMCRLVTDAAQNGNEQVIKDVRKITRSGDDWIPDSPQDLASKIFHS
CFMGTENSSKETRNRAKDLSNAIGSYHVDLKMDSLVSSVVSLFEVATGKKPIYKIFGGSQ
IENLALQNIQARLRMVLSYLFAQLLPWVRGIPNSGGLLVLGSANVDECLRGYLTKYDCSS
ADINPIGGISKTDLKRFIAYASKQYNMPILNDFLNATPTAELEPMTKDYVQSDEIDMGMT
YEELGVFGYLRKVEKCGPYSMFLKLLHQWSPKLTPRQISEKVKRFFFFYAINRHKQTVLT
PSYHAEQYSPEDNRFDLRPFLINPRFPWASRKIDEVVEQCEAHKGSTLDIMSID |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Johnston, M., Andrews, S., Brinkman, R., Cooper, J., Ding, H., Dover, J., Du, Z., Favello, A., Fulton, L., Gattung, S., et, a. l. .. (1994). "Complete nucleotide sequence of Saccharomyces cerevisiae chromosome VIII." Science 265:2077-2082.8091229
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
|
|---|
