D-3-phosphoglycerate dehydrogenase 2 (P40510)
| Identification | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | D-3-phosphoglycerate dehydrogenase 2 | ||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||
| Gene Name | SER33 | ||||||||||||||||||||||||||||||
| Enzyme Class | |||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||
| General Function | Involved in oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor | ||||||||||||||||||||||||||||||
| Specific Function | 3-phospho-D-glycerate + NAD(+) = 3- phosphonooxypyruvate + NADH | ||||||||||||||||||||||||||||||
| Cellular Location | Not Available | ||||||||||||||||||||||||||||||
| SMPDB Pathways |
| ||||||||||||||||||||||||||||||
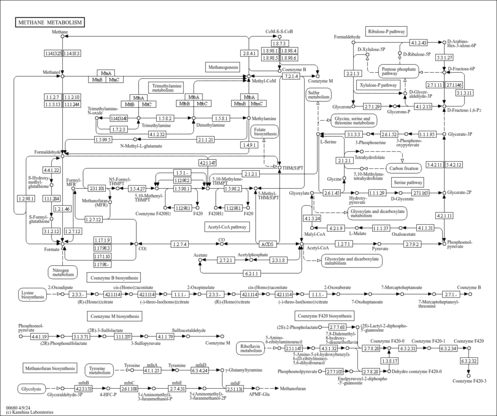
| KEGG Pathways |
| ||||||||||||||||||||||||||||||
| SMPDB Reactions |
| ||||||||||||||||||||||||||||||
| KEGG Reactions |
| ||||||||||||||||||||||||||||||
| Metabolites |
| ||||||||||||||||||||||||||||||
| GO Classification |
| ||||||||||||||||||||||||||||||
| Gene Properties | |||||||||||||||||||||||||||||||
| Chromosome Location | chromosome 9 | ||||||||||||||||||||||||||||||
| Locus | YIL074C | ||||||||||||||||||||||||||||||
| Gene Sequence | >1410 bp ATGTCTTATTCAGCTGCCGATAATTTACAAGATTCATTCCAACGTGCCATGAACTTTTCT GGCTCTCCTGGTGCAGTCTCAACCTCACCAACTCAGTCATTTATGAACACACTACCTCGT CGTGTAAGCATTACAAAGCAACCAAAGGCTTTAAAACCTTTTTCTACTGGTGACATGAAT ATTCTACTGTTGGAAAATGTCAATGCAACTGCAATCAAAATCTTCAAGGATCAGGGTTAC CAAGTAGAGTTCCACAAGTCTTCTCTACCTGAGGATGAATTGATTGAAAAAATCAAAGAC GTACACGCTATCGGTATAAGATCCAAAACTAGATTGACTGAAAAAATACTACAGCATGCC AGGAATCTAGTTTGTATTGGTTGTTTTTGCATAGGTACCAATCAAGTAGACCTAAAATAT GCCGCTAGTAAAGGTATTGCTGTTTTCAATTCGCCATTCTCCAATTCAAGATCCGTAGCA GAATTGGTAATTGGTGAGATCATTAGTTTAGCAAGACAATTAGGTGATAGATCCATTGAA CTGCATACAGGTACATGGAATAAAGTCGCTGCTAGGTGTTGGGAAGTAAGAGGAAAAACT CTCGGTATTATTGGGTATGGTCACATTGGTTCGCAATTATCAGTTCTTGCAGAAGCTATG GGCCTGCATGTGCTATACTATGATATCGTGACAATTATGGCCTTAGGTACTGCCAGACAA GTTTCTACATTAGATGAATTGTTGAATAAATCTGATTTTGTAACACTACATGTACCAGCT ACTCCAGAAACTGAAAAAATGTTATCTGCTCCACAATTCGCTGCTATGAAGGACGGGGCT TATGTTATTAATGCCTCAAGAGGTACTGTCGTGGACATTCCATCTCTGATCCAAGCCGTC AAGGCCAACAAAATTGCAGGTGCTGCTTTAGATGTTTATCCACATGAACCAGCTAAGAAC GGTGAAGGTTCATTTAACGATGAACTTAACAGCTGGACTTCTGAGTTGGTTTCATTACCA AATATAATCCTGACACCACATATTGGTGGCTCTACAGAAGAAGCTCAAAGTTCAATCGGT ATTGAGGTGGCTACTGCATTGTCCAAATACATCAATGAAGGTAACTCTGTCGGTTCTGTG AACTTCCCAGAAGTCAGTTTGAAGTCTTTGGACTACGATCAAGAGAACACAGTACGTGTC TTGTATATTCATCGTAACGTTCCTGGTGTTTTGAAGACCGTTAATGATATCTTATCCGAT CATAATATCGAGAAACAGTTTTCTGATTCTCACGGCGAGATCGCTTATCTAATGGCAGAC ATCTCTTCTGTTAATCAAAGTGAAATCAAGGATATATATGAAAAGTTGAACCAAACTTCT GCCAAAGTTTCCATCAGGTTATTATACTAA | ||||||||||||||||||||||||||||||
| Protein Properties | |||||||||||||||||||||||||||||||
| Pfam Domain Function | |||||||||||||||||||||||||||||||
| Protein Residues | 469 | ||||||||||||||||||||||||||||||
| Protein Molecular Weight | 51187.80078 | ||||||||||||||||||||||||||||||
| Protein Theoretical pI | 6.38 | ||||||||||||||||||||||||||||||
| Signalling Regions |
| ||||||||||||||||||||||||||||||
| Transmembrane Regions |
| ||||||||||||||||||||||||||||||
| Protein Sequence | >D-3-phosphoglycerate dehydrogenase 2 MSYSAADNLQDSFQRAMNFSGSPGAVSTSPTQSFMNTLPRRVSITKQPKALKPFSTGDMN ILLLENVNATAIKIFKDQGYQVEFHKSSLPEDELIEKIKDVHAIGIRSKTRLTEKILQHA RNLVCIGCFCIGTNQVDLKYAASKGIAVFNSPFSNSRSVAELVIGEIISLARQLGDRSIE LHTGTWNKVAARCWEVRGKTLGIIGYGHIGSQLSVLAEAMGLHVLYYDIVTIMALGTARQ VSTLDELLNKSDFVTLHVPATPETEKMLSAPQFAAMKDGAYVINASRGTVVDIPSLIQAV KANKIAGAALDVYPHEPAKNGEGSFNDELNSWTSELVSLPNIILTPHIGGSTEEAQSSIG IEVATALSKYINEGNSVGSVNFPEVSLKSLDYDQENTVRVLYIHRNVPGVLKTVNDILSD HNIEKQFSDSHGEIAYLMADISSVNQSEIKDIYEKLNQTSAKVSIRLLY | ||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||
| External Links |
| ||||||||||||||||||||||||||||||
| General Reference |
| ||||||||||||||||||||||||||||||