| Identification |
|---|
| Name | Succinate dehydrogenase [ubiquinone] flavoprotein subunit 2, mitochondrial |
|---|
| Synonyms | - Flavoprotein subunit of complex II
- FP
- SDH1b
|
|---|
| Gene Name | Not Available |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in electron carrier activity |
|---|
| Specific Function | Probable minor catalytic subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q). Probably forms a catalytic dimer with SDH2. Electrons flow from succinate to the FAD bound to the catalytic subunit, and sequentially through the iron-sulfur clusters bound to SDH2 and enter the membrane dimer formed by SDH3 and SDH4 |
|---|
| Cellular Location | Mitochondrion inner membrane; Peripheral membrane protein; Matrix side |
|---|
| SMPDB Pathways | |
|---|
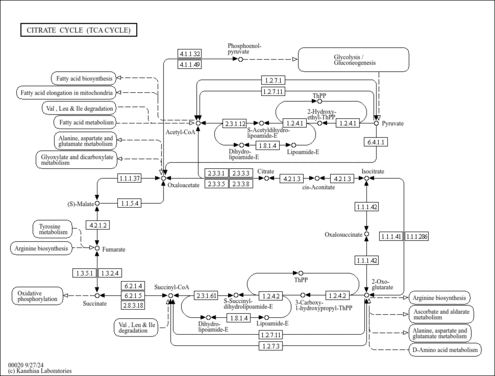
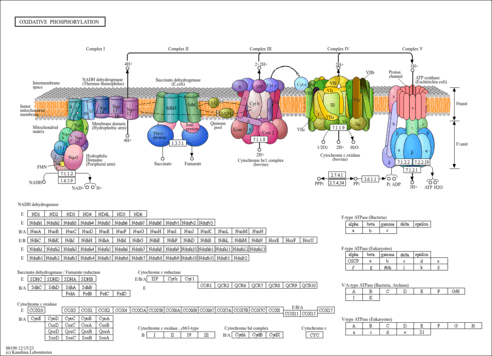
| KEGG Pathways | |
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | Not Available |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00088 | Ubiquinone Q1 | Show | | YMDB00101 | Fumaric acid | Show | | YMDB00338 | Succinic acid | Show | | YMDB00902 | ubiquinol | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| binding | | nucleoside binding | | purine nucleoside binding | | adenyl nucleotide binding | | FAD or FADH2 binding | | oxidoreductase activity | | electron carrier activity | | catalytic activity | | oxidoreductase activity, acting on the CH-CH group of donors | | Process |
|---|
| oxidation reduction | | cofactor metabolic process | | coenzyme metabolic process | | acetyl-CoA metabolic process | | acetyl-CoA catabolic process | | tricarboxylic acid cycle | | metabolic process | | generation of precursor metabolites and energy | | cellular metabolic process | | electron transport chain |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | Not Available |
|---|
| Locus | YJL045W |
|---|
| Gene Sequence | >1905 bp
ATGTTATCTTTGAAAAAAGGAATAACAAAATCATACATCTTGCAAAGAACTTTCACTTCT
TCCTCTGTTGTTCGTCAAATTGGGGAAGTGAAATCTGAATCGAAACCGCCGGCCAAATAT
CATATTATCGACCATGAATATGATTGTGTGGTGGTAGGCGCTGGCGGTGCAGGTTTAAGA
GCAGCTTTCGGTTTGGCTGAAGCTGGATACAAGACTGCTTGTTTATCCAAGTTGTTTCCA
ACAAGGTCACATACTGTGGCTGCTCAGGGTGGAATTAATGCTGCGCTGGGAAATATGCAT
CCAGATGATTGGAAATCGCACATGTACGACACTGTCAAGGGTTCTGACTGGCTCGGAGAC
CAAGATGCAATCCATTACATGACAAGAGAAGCACCTAAGTCTGTCATTGAACTAGAACAT
TACGGTATGCCCTTTTCGAGGACTGAAGATGGAAGGATTTACCAGAGAGCATTTGGGGGA
CAATCCAAAGATTTTGGTAAAGGTGGACAGGCCTATAGGACTTGTGCGGTGGCAGATAGA
ACAGGTCACGCAATGCTTCATACATTGTATGGACAAGCGCTGAAAAATAATACACACTTC
TTTATTGAATACTTTGCAATGGATTTGTTGACCCATAATGGCGAGGTTGTGGGTGTCATT
GCCTATAATCAGGAGGACGGTACAATTCACAGATTCAGAGCACATAAGACCGTCATCGCG
ACAGGCGGATACGGTAGAGCTTACTTCTCTTGCACTTCTGCTCACACTTGTACAGGTGAC
GGTAATGCTATGGTTTCTCGCGCTGGATTTCCACTAGAGGATTTAGAATTTGTTCAATTT
CATCCGTCAGGAATTTATGGGTCTGGCTGCCTAATCACTGAAGGTGCCCGTGGTGAGGGT
GGATTTTTATTGAATTCTGAAGGAGAAAGGTTTATGGAACGCTATGCTCCTACTGCCAAG
GACTTGGCAAGCAGGGATGTTGTTTCCAGAGCAATCACCATGGAAATCAGGGCTGGCAGA
GGTGTCGGGAAAAACAAGGATCATATCCTTTTACAATTAAGCCATCTACCACCTGAGGTA
CTAAAGGAAAGGCTACCGGGAATATCTGAAACAGCTGCTGTCTTTGCGGGTGTCGATGTC
ACCCAGGAGCCAATTCCTGTCTTGCCAACTGTCCATTATAATATGGGAGGCATTCCCACA
AAATGGACTGGTGAAGCATTGACCATTGACGAGGAAACTGGAGAGGATAAGGTCATCCCA
GGATTGATGGCGTGTGGTGAAGCTGCTTGCGTATCGGTTCATGGAGCGAACAGATTAGGC
GCTAACTCACTACTGGATTTAGTCGTTTTCGGTCGCGCCGTTGCAAATACCATTGCTGAC
ACATTACAGCCTGGCTTGCCTCATAAGCCATTGGCTTCAAACATCGGGCACGAGTCAATT
GCTAATTTGGATAAAGTAAGAAATGCTCGCGGCTCACTGAAAACCTCTCAAATCAGGTTG
AACATGCAAAGGACAATGCAAAAAGATGTTTCTGTTTTCAGGACGCAAGACACTCTAGAT
GAAGGTGTTAGAAATATTACTGAAGTGGACAAGACATTTGAGGATGTGCACGTTTCTGAT
AAGTCAATGATCTGGAATTCTGATCTCGTAGAAACTCTGGAATTGCAAAATTTACTTACT
TGTGCCACACAAACGGCTGTTTCTGCTTCCAAAAGAAAGGAGTCTCGTGGTGCTCATGCG
AGAGAGGACTATGCAAAAAGAGATGATGTGAATTGGAGAAAGCACACATTATCATGGCAA
AAGGGGACATCAACACCTGTAAAAATCAAGTACAGGAATGTAATCGCACATACTTTAGAT
GAGAATGAATGCGCCCCAGTCCCTCCAGCTGTCAGATCCTATTAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 634 |
|---|
| Protein Molecular Weight | 69382.0 |
|---|
| Protein Theoretical pI | 7.4 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Succinate dehydrogenase [ubiquinone] flavoprotein subunit 2, mitochondrial
MLSLKKGITKSYILQRTFTSSSVVRQIGEVKSESKPPAKYHIIDHEYDCVVVGAGGAGLR
AAFGLAEAGYKTACLSKLFPTRSHTVAAQGGINAALGNMHPDDWKSHMYDTVKGSDWLGD
QDAIHYMTREAPKSVIELEHYGMPFSRTEDGRIYQRAFGGQSKDFGKGGQAYRTCAVADR
TGHAMLHTLYGQALKNNTHFFIEYFAMDLLTHNGEVVGVIAYNQEDGTIHRFRAHKTVIA
TGGYGRAYFSCTSAHTCTGDGNAMVSRAGFPLEDLEFVQFHPSGIYGSGCLITEGARGEG
GFLLNSEGERFMERYAPTAKDLASRDVVSRAITMEIRAGRGVGKNKDHILLQLSHLPPEV
LKERLPGISETAAVFAGVDVTQEPIPVLPTVHYNMGGIPTKWTGEALTIDEETGEDKVIP
GLMACGEAACVSVHGANRLGANSLLDLVVFGRAVANTIADTLQPGLPHKPLASNIGHESI
ANLDKVRNARGSLKTSQIRLNMQRTMQKDVSVFRTQDTLDEGVRNITEVDKTFEDVHVSD
KSMIWNSDLVETLELQNLLTCATQTAVSASKRKESRGAHAREDYAKRDDVNWRKHTLSWQ
KGTSTPVKIKYRNVIAHTLDENECAPVPPAVRSY |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Galibert, F., Alexandraki, D., Baur, A., Boles, E., Chalwatzis, N., Chuat, J. C., Coster, F., Cziepluch, C., De Haan, M., Domdey, H., Durand, P., Entian, K. D., Gatius, M., Goffeau, A., Grivell, L. A., Hennemann, A., Herbert, C. J., Heumann, K., Hilger, F., Hollenberg, C. P., Huang, M. E., Jacq, C., Jauniaux, J. C., Katsoulou, C., Karpfinger-Hartl, L., et, a. l. .. (1996). "Complete nucleotide sequence of Saccharomyces cerevisiae chromosome X." EMBO J 15:2031-2049.8641269
- Colby, G., Ishii, Y., Tzagoloff, A. (1998). "Suppression of sdh1 mutations by the SDH1b gene of Saccharomyces cerevisiae." Yeast 14:1001-1006.9730279
|
|---|