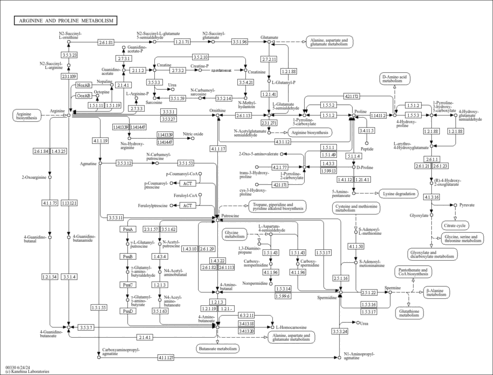
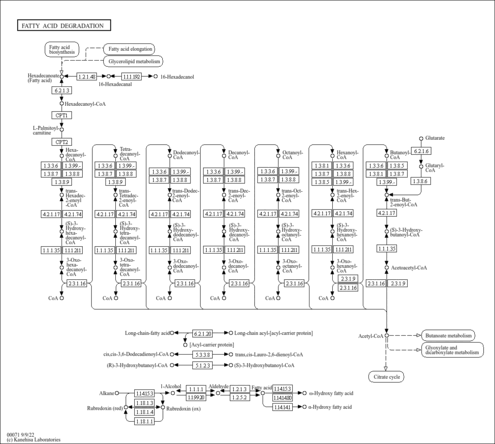
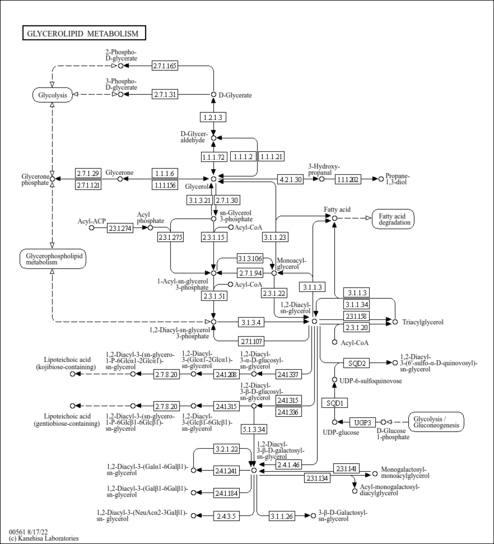
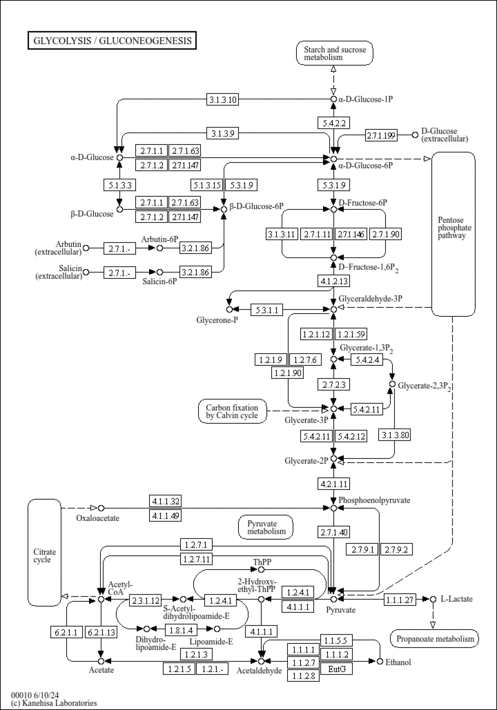
| General Reference | - Dujon, B., Albermann, K., Aldea, M., Alexandraki, D., Ansorge, W., Arino, J., Benes, V., Bohn, C., Bolotin-Fukuhara, M., Bordonne, R., Boyer, J., Camasses, A., Casamayor, A., Casas, C., Cheret, G., Cziepluch, C., Daignan-Fornier, B., Dang, D. V., de Haan, M., Delius, H., Durand, P., Fairhead, C., Feldmann, H., Gaillon, L., Kleine, K., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome XV." Nature 387:98-102.9169874
- Chalmers, R. M., Keen, J. N., Fewson, C. A. (1991). "Comparison of benzyl alcohol dehydrogenases and benzaldehyde dehydrogenases from the benzyl alcohol and mandelate pathways in Acinetobacter calcoaceticus and from the TOL-plasmid-encoded toluene pathway in Pseudomonas putida. N-terminal amino acid sequences, amino acid compositions and immunological cross-reactions." Biochem J 273(Pt 1):99-107.1989592
- Larsson, T., Norbeck, J., Karlsson, H., Karlsson, K. A., Blomberg, A. (1997). "Identification of two-dimensional gel electrophoresis resolved yeast proteins by matrix-assisted laser desorption ionization mass spectrometry." Electrophoresis 18:418-423.9150920
- Tessier, W. D., Meaden, P. G., Dickinson, F. M., Midgley, M. (1998). "Identification and disruption of the gene encoding the K(+)-activated acetaldehyde dehydrogenase of Saccharomyces cerevisiae." FEMS Microbiol Lett 164:29-34.9675847
- Grandier-Vazeille, X., Bathany, K., Chaignepain, S., Camougrand, N., Manon, S., Schmitter, J. M. (2001). "Yeast mitochondrial dehydrogenases are associated in a supramolecular complex." Biochemistry 40:9758-9769.11502169
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
- Reinders, J., Wagner, K., Zahedi, R. P., Stojanovski, D., Eyrich, B., van der Laan, M., Rehling, P., Sickmann, A., Pfanner, N., Meisinger, C. (2007). "Profiling phosphoproteins of yeast mitochondria reveals a role of phosphorylation in assembly of the ATP synthase." Mol Cell Proteomics 6:1896-1906.17761666
|
|---|