| Identification |
|---|
| Name | NAD-dependent malic enzyme, mitochondrial |
|---|
| Synonyms | |
|---|
| Gene Name | MAE1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in oxidoreductase activity |
|---|
| Specific Function | (S)-malate + NAD(+) = pyruvate + CO(2) + NADH |
|---|
| Cellular Location | Mitochondrion matrix |
|---|
| SMPDB Pathways | |
|---|
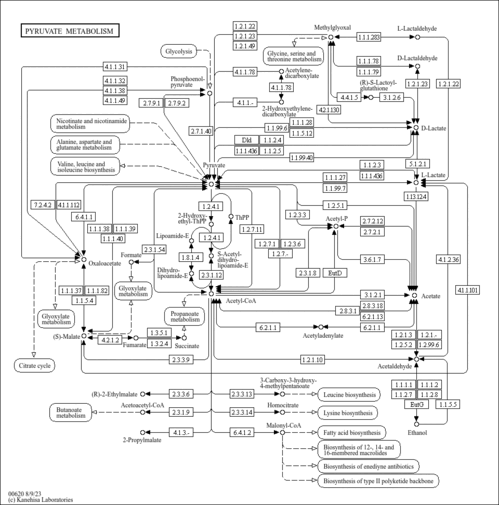
| KEGG Pathways | |
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00110 | NAD | Show | | YMDB00143 | NADH | Show | | YMDB00175 | Pyruvic acid | Show | | YMDB00235 | Oxalacetic acid | Show | | YMDB00280 | (S)-Malic acid | Show | | YMDB00412 | NAD(+) | Show | | YMDB00426 | NADPH | Show | | YMDB00427 | NADP | Show | | YMDB00862 | hydron | Show | | YMDB00912 | Carbon dioxide | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| oxidoreductase activity | | oxidoreductase activity, acting on CH-OH group of donors | | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor | | catalytic activity | | NAD or NADH binding | | malate dehydrogenase activity | | binding | | malic enzyme activity | | nucleotide binding | | ion binding | | cation binding | | metal ion binding | | Process |
|---|
| oxidation reduction | | organic acid metabolic process | | oxoacid metabolic process | | carboxylic acid metabolic process | | metabolic process | | dicarboxylic acid metabolic process | | cellular metabolic process | | malate metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 11 |
|---|
| Locus | YKL029C |
|---|
| Gene Sequence | >2010 bp
ATGCTTAGAACCAGACTATCCGTTTCCGTTGCTGCTAGATCGCAACTAACCAGATCCTTG
ACAGCATCAAGGACAGCACCATTAAGAAGATGGCCTATTCAGCAATCGCGTTTATATTCT
TCTAACACTAGATCGCATAAAGCTACCACAACAAGAGAAAATACTTTCCAAAAGCCATAC
AGCGACGAGGAGGTCACTAAAACACCCGTCGGTTCTCGCGCCAGAAAGATCTTCGAAGCT
CCTCACCCACATGCCACTCGTTTGACTGTAGAAGGTGCCATAGAATGTCCCTTGGAGAGC
TTTCAACTTTTAAACTCTCCTTTATTTAACAAGGGTTCTGCATTTACACAAGAAGAAAGG
GAAGCGTTTAATTTAGAAGCATTGCTACCACCACAAGTGAACACTTTGGACGAACAACTG
GAAAGAAGCTACAAGCAGTTATGCTATTTGAAGACGCCCTTGGCCAAAAACGACTTCATG
ACGTCTTTGAGAGTACAGAACAAAGTCCTATATTTTGCATTAATAAGGAGACATATCAAG
GAATTAGTTCCTATCATTTACACCCCAACCGAAGGTGATGCTATTGCTGCCTATTCCCAC
AGGTTCAGAAAGCCAGAAGGTGTGTTTTTAGACATTACCGAACCTGATTCCATCGAATGT
AGATTGGCTACATACGGTGGAGACAAAGATGTAGACTACATCGTTGTGTCGGATTCGGAA
GGTATTCTGGGAATTGGTGACCAAGGTATCGGTGGTGTACGTATTGCTATCTCCAAATTG
GCATTGATGACGCTGTGCGGTGGTATTCATCCCGGCCGTGTGCTACCTGTGTGTTTGGAC
GTCGGTACTAACAACAAGAAACTAGCCCGTGACGAATTGTACATGGGTAACAAGTTCTCC
AGAATCAGGGGTAAGCAATATGACGACTTCTTGGAAAAATTCATCAAGGCCGTTAAGAAA
GTGTATCCAAGCGCCGTTCTGCATTTCGAAGATTTCGGTGTTAAGAACGCTAGAAGATTA
CTAGAAAAGTACAGGTACGAATTGCCATCATTCAACGATGACATTCAGGGCACCGGTGCC
GTCGTGATGGCCTCGTTGATTGCTGCTTTGAAACATACCAACAGAGACTTGAAAGACACC
AGAGTGCTTATTTACGGTGCCGGGTCTGCGGGCCTCGGTATCGCAGATCAAATTGTGAAT
CATATGGTCACGCACGGCGTTGACAAGGAAGAAGCGCGCAAGAAAATCTTCTTGATGGAC
AGACGTGGGTTAATTCTACAATCTTACGAGGCTAACTCCACTCCCGCCCAACACGTATAC
GCTAAGAGTGATGCGGAATGGGCTGGTATCAACACCCGCTCTTTACATGATGTGGTGGAG
AACGTCAAACCAACGTGTTTGGTTGGCTGCTCCACACAAGCAGGCGCATTCACTCAAGAT
GTCGTAGAAGAAATGCACAAGCACAATCCTAGACCGATCATTTTCCCATTATCCAACCCT
ACTAGACTACACGAAGCCGTTCCTGCCGATTTAATGAAGTGGACCAACAACAACGCTCTT
GTAGCTACCGGATCTCCTTTCCCACCTGTTGATGGTTACCGTATCTCGGAGAACAACAAT
TGTTACTCTTTCCCAGGTATCGGTTTAGGTGCCGTACTATCGCGTGCCACCACCATCACA
GACAAGATGATCTCCGCTGCAGTGGACCAACTAGCCGAATTGTCGCCACTAAGAGAGGGC
GACTCGAGACCTGGGTTGCTACCCGGCCTGGACACCATCACCAACACTTCTGCGCGTCTA
GCTACCGCTGTGATCTTGCAAGCACTCGAGGAGGGAACCGCCCGTATCGAGCAAGAACAA
GTACCGGGAGGAGCTCCCGGCGAAACTGTCAAGGTTCCTCGTGACTTTGACGAATGTTTA
CAGTGGGTCAAAGCCCAAATGTGGGAGCCTGTGTACAGACCTATGATCAAGGTCCAACAT
GACCCATCGGTGCACACCAACCAATTGTAG |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 669 |
|---|
| Protein Molecular Weight | 74375.29688 |
|---|
| Protein Theoretical pI | 8.3 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >NAD-dependent malic enzyme, mitochondrial
MLRTRLSVSVAARSQLTRSLTASRTAPLRRWPIQQSRLYSSNTRSHKATTTRENTFQKPY
SDEEVTKTPVGSRARKIFEAPHPHATRLTVEGAIECPLESFQLLNSPLFNKGSAFTQEER
EAFNLEALLPPQVNTLDEQLERSYKQLCYLKTPLAKNDFMTSLRVQNKVLYFALIRRHIK
ELVPIIYTPTEGDAIAAYSHRFRKPEGVFLDITEPDSIECRLATYGGDKDVDYIVVSDSE
GILGIGDQGIGGVRIAISKLALMTLCGGIHPGRVLPVCLDVGTNNKKLARDELYMGNKFS
RIRGKQYDDFLEKFIKAVKKVYPSAVLHFEDFGVKNARRLLEKYRYELPSFNDDIQGTGA
VVMASLIAALKHTNRDLKDTRVLIYGAGSAGLGIADQIVNHMVTHGVDKEEARKKIFLMD
RRGLILQSYEANSTPAQHVYAKSDAEWAGINTRSLHDVVENVKPTCLVGCSTQAGAFTQD
VVEEMHKHNPRPIIFPLSNPTRLHEAVPADLMKWTNNNALVATGSPFPPVDGYRISENNN
CYSFPGIGLGAVLSRATTITDKMISAAVDQLAELSPLREGDSRPGLLPGLDTITNTSARL
ATAVILQALEEGTARIEQEQVPGGAPGETVKVPRDFDECLQWVKAQMWEPVYRPMIKVQH
DPSVHTNQL |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Dujon, B., Alexandraki, D., Andre, B., Ansorge, W., Baladron, V., Ballesta, J. P., Banrevi, A., Bolle, P. A., Bolotin-Fukuhara, M., Bossier, P., et, a. l. .. (1994). "Complete DNA sequence of yeast chromosome XI." Nature 369:371-378.8196765
- Boles, E., de Jong-Gubbels, P., Pronk, J. T. (1998). "Identification and characterization of MAE1, the Saccharomyces cerevisiae structural gene encoding mitochondrial malic enzyme." J Bacteriol 180:2875-2882.9603875
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
- Sickmann, A., Reinders, J., Wagner, Y., Joppich, C., Zahedi, R., Meyer, H. E., Schonfisch, B., Perschil, I., Chacinska, A., Guiard, B., Rehling, P., Pfanner, N., Meisinger, C. (2003). "The proteome of Saccharomyces cerevisiae mitochondria." Proc Natl Acad Sci U S A 100:13207-13212.14576278
|
|---|