| Identification |
|---|
| Name | Acid phosphatase PHO11 |
|---|
| Synonyms | |
|---|
| Gene Name | PHO11 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in acid phosphatase activity |
|---|
| Specific Function | A phosphate monoester + H(2)O = an alcohol + phosphate |
|---|
| Cellular Location | Cytoplasmic |
|---|
| SMPDB Pathways | |
|---|
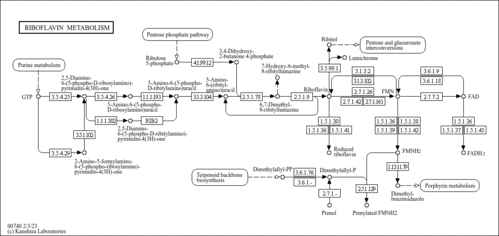
| KEGG Pathways | |
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00085 | Flavin Mononucleotide | Show | | YMDB00164 | Thiamine monophosphate | Show | | YMDB00220 | Thiamine | Show | | YMDB00534 | Riboflavin | Show | | YMDB00890 | water | Show | | YMDB00907 | phosphate | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| catalytic activity | | hydrolase activity | | hydrolase activity, acting on ester bonds | | phosphoric ester hydrolase activity | | phosphatase activity | | acid phosphatase activity | | Process |
|---|
| Not Available |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 1 |
|---|
| Locus | YAR071W |
|---|
| Gene Sequence | >1404 bp
ATGTTGAAGTCAGCCGTTTATTCAATTTTAGCCGCTTCTTTGGTTAATGCAGGTACCATA
CCCCTCGGAAAGTTATCTGACATTGACAAAATCGGAACTCAAACGGAAATTTTCCCATTT
TTGGGTGGTTCTGGGCCATACTACTCTTTCCCTGGTGATTATGGTATTTCTCGTGATTTG
CCGGAAAGTTGTGAAATGAAGCAAGTGCAAATGGTTGGTAGACACGGTGAAAGATACCCC
ACTGTCAGCAAAGCCAAAAGTATCATGACAACATGGTACAAATTGAGTAACTATACCGGT
CAATTCAGCGGAGCATTGTCTTTCTTGAACGATGACTACGAATTTTTCATTCGTGACACC
AAAAACCTAGAAATGGAAACCACACTTGCCAATTCGGTCAATGTTTTGAACCCATATACC
GGTGAGATGAATGCTAAGAGACACGCTCGTGATTTCTTGGCGCAATATGGCTACATGGTC
GAAAACCAAACCAGTTTTGCCGTTTTTACGTCTAACTCGAACAGATGTCATGATACTGCC
CAGTATTTCATTGACGGTTTGGGTGATAAATTCAACATATCCTTGCAAACCATCAGTGAA
GCCGAGTCTGCTGGTGCCAATACTCTGAGTGCCCACCATTCGTGTCCTGCTTGGGACGAT
GATGTCAACGATGACATTTTGAAAAAATATGATACCAAATATTTGAGTGGTATTGCCAAG
AGATTAAACAAGGAAAACAAGGGTTTGAATCTGACTTCAAGTGATGCAAACACTTTTTTT
GCATGGTGTGCATATGAAATAAACGCTAGAGGTTACAGTGACATCTGTAACATCTTCACC
AAAGATGAATTGGTCCGTTTCTCCTACGGCCAAGACTTGGAAACTTATTATCAAACGGGA
CCAGGCTATGACGTCGTCAGATCCGTCGGTGCCAACTTGTTCAACGCTTCAGTGAAACTA
CTAAAGGAAAGTGAGGTCCAGGACCAAAAGGTTTGGTTGAGTTTCACCCACGATACCGAT
ATTCTGAACTATTTGACCACTATCGGCATAATCGATGACAAAAATAACTTGACCGCCGAA
CATGTTCCATTCATGGAAAACACTTTCCACAGATCCTGGTACGTTCCACAAGGTGCTCGT
GTTTACACTGAAAAGTTCCAGTGTTCCAATGACACCTATGTTAGATACGTCATCAACGAT
GCTGTCGTTCCAATTGAAACCTGTTCTACTGGTCCAGGGTTCTCCTGTGAAATAAATGAC
TTCTACGACTATGCTGAAAAGAGAGTAGCCGGTACTGACTTCCTAAAGGTCTGTAACGTC
AGCAGCGTCAGTAACTCTACTGAATTGACCTTTTTCTGGGACTGGAATACCAAGCACTAC
AACGACACTTTATTAAAACAGTAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 467 |
|---|
| Protein Molecular Weight | 52757.39844 |
|---|
| Protein Theoretical pI | 4.78 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Acid phosphatase PHO11
MLKSAVYSILAASLVNAGTIPLGKLSDIDKIGTQTEIFPFLGGSGPYYSFPGDYGISRDL
PESCEMKQVQMVGRHGERYPTVSKAKSIMTTWYKLSNYTGQFSGALSFLNDDYEFFIRDT
KNLEMETTLANSVNVLNPYTGEMNAKRHARDFLAQYGYMVENQTSFAVFTSNSNRCHDTA
QYFIDGLGDKFNISLQTISEAESAGANTLSAHHSCPAWDDDVNDDILKKYDTKYLSGIAK
RLNKENKGLNLTSSDANTFFAWCAYEINARGYSDICNIFTKDELVRFSYGQDLETYYQTG
PGYDVVRSVGANLFNASVKLLKESEVQDQKVWLSFTHDTDILNYLTTIGIIDDKNNLTAE
HVPFMENTFHRSWYVPQGARVYTEKFQCSNDTYVRYVINDAVVPIETCSTGPGFSCEIND
FYDYAEKRVAGTDFLKVCNVSSVSNSTELTFFWDWNTKHYNDTLLKQ |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Bussey, H., Kaback, D. B., Zhong, W., Vo, D. T., Clark, M. W., Fortin, N., Hall, J., Ouellette, B. F., Keng, T., Barton, A. B., et, a. l. .. (1995). "The nucleotide sequence of chromosome I from Saccharomyces cerevisiae." Proc Natl Acad Sci U S A 92:3809-3813.7731988
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
|
|---|