Pyruvate carboxylase 2 (P32327)
| Identification | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Pyruvate carboxylase 2 | |||||||||||||||||||||||||||
| Synonyms |
| |||||||||||||||||||||||||||
| Gene Name | PYC2 | |||||||||||||||||||||||||||
| Enzyme Class | ||||||||||||||||||||||||||||
| Biological Properties | ||||||||||||||||||||||||||||
| General Function | Involved in catalytic activity | |||||||||||||||||||||||||||
| Specific Function | Pyruvate carboxylase catalyzes a 2-step reaction, involving the ATP-dependent carboxylation of the covalently attached biotin in the first step and the transfer of the carboxyl group to pyruvate in the second | |||||||||||||||||||||||||||
| Cellular Location | Cytoplasm | |||||||||||||||||||||||||||
| SMPDB Pathways |
| |||||||||||||||||||||||||||
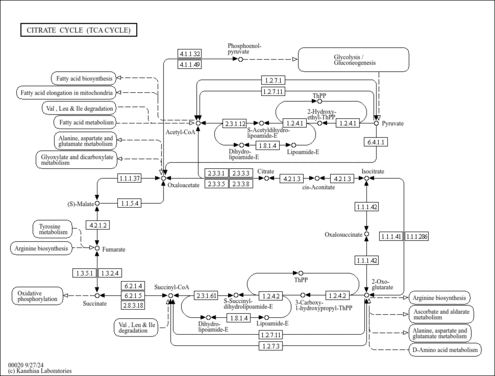
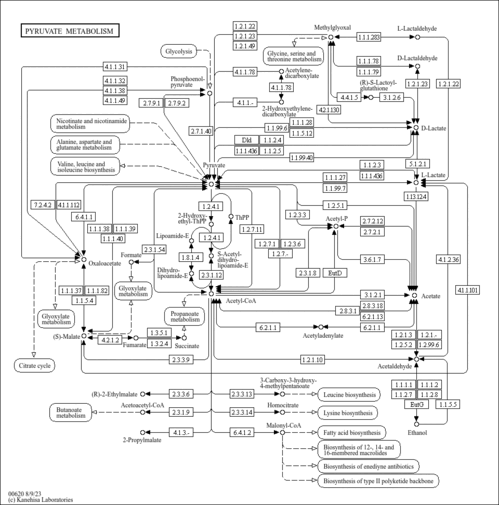
| KEGG Pathways |
| |||||||||||||||||||||||||||
| SMPDB Reactions |
| |||||||||||||||||||||||||||
| KEGG Reactions |
| |||||||||||||||||||||||||||
| Metabolites |
| |||||||||||||||||||||||||||
| GO Classification |
| |||||||||||||||||||||||||||
| Gene Properties | ||||||||||||||||||||||||||||
| Chromosome Location | chromosome 2 | |||||||||||||||||||||||||||
| Locus | YBR218C | |||||||||||||||||||||||||||
| Gene Sequence | >3543 bp ATGAGCAGTAGCAAGAAATTGGCCGGTCTTAGGGACAATTTCAGTTTGCTCGGCGAAAAG AATAAGATCTTGGTCGCCAATAGAGGTGAAATTCCGATTAGAATTTTTAGATCTGCTCAT GAGCTGTCTATGAGAACCATCGCCATATACTCCCATGAGGACCGTCTTTCAATGCACAGG TTGAAGGCGGACGAAGCGTATGTTATCGGGGAGGAGGGCCAGTATACACCTGTGGGTGCT TACTTGGCAATGGACGAGATCATCGAAATTGCAAAGAAGCATAAGGTGGATTTCATCCAT CCAGGTTATGGGTTCTTGTCTGAAAATTCGGAATTTGCCGACAAAGTAGTGAAGGCCGGT ATCACTTGGATCGGCCCTCCAGCTGAAGTTATTGACTCTGTGGGTGACAAAGTCTCTGCC AGACACTTGGCAGCAAGAGCTAACGTTCCTACCGTTCCCGGTACTCCAGGACCTATCGAA ACTGTGCAAGAGGCACTTGACTTCGTTAATGAATACGGCTACCCGGTGATCATTAAGGCC GCCTTTGGTGGTGGTGGTAGAGGTATGAGAGTCGTTAGAGAAGGTGACGACGTGGCAGAT GCCTTTCAACGTGCTACCTCCGAAGCCCGTACTGCCTTCGGTAATGGTACCTGCTTTGTG GAAAGATTCTTGGACAAGCCAAAGCATATTGAAGTTCAATTGTTGGCTGATAACCACGGA AACGTGGTTCATCTTTTCGAAAGAGACTGTTCTGTGCAAAGAAGACACCAAAAAGTTGTC GAAGTCGCTCCAGCAAAGACTTTGCCCCGTGAAGTTCGTGACGCTATTTTGACAGATGCT GTTAAATTAGCTAAGGTATGTGGTTACAGAAACGCAGGTACCGCCGAATTCTTGGTTGAC AACCAAAACAGACACTATTTCATTGAAATTAATCCAAGAATTCAAGTGGAGCATACCATC ACTGAAGAAATCACCGGTATTGACATTGTTTCTGCCCAAATCCAGATTGCCGCAGGTGCC ACTTTGACTCAACTAGGTCTATTACAGGATAAAATCACCACCCGTGGGTTTTCCATCCAA TGTCGTATTACCACTGAAGATCCCTCTAAGAATTTCCAACCGGATACCGGTCGCCTGGAG GTCTATCGTTCTGCCGGTGGTAATGGTGTGAGATTGGACGGTGGTAACGCTTATGCAGGT GCTACTATCTCGCCTCACTACGACTCAATGCTGGTCAAATGTTCATGCTCTGGTTCTACT TATGAAATCGTCCGTAGGAAGATGATTCGTGCCCTGATCGAATTCAGAATCAGAGGTGTT AAGACCAACATTCCCTTCCTATTGACTCTTTTGACCAATCCAGTTTTTATTGAGGGTACA TACTGGACGACTTTTATTGACGACACCCCACAACTGTTCCAAATGGTATCGTCACAAAAC AGAGCGCAAAAACTGTTACACTATTTGGCAGACTTGGCAGTTAACGGTTCTTCTATTAAG GGTCAAATTGGCTTGCCAAAACTAAAATCAAATCCAAGTGTCCCCCATTTGCACGATGCT CAGGGCAATGTCATCAACGTTACAAAGTCTGCACCACCATCCGGATGGAGACAAGTGCTA CTGGAAAAGGGACCATCTGAATTTGCCAAGCAAGTCAGACAGTTCAATGGTACTCTACTG ATGGACACCACCTGGAGAGACGCTCATCAATCTCTACTTGCAACAAGAGTCAGAACCCAC GATTTGGCTACAATCGCTCCAACAACCGCACATGCCCTTGCAGGTGCTTTCGCTTTAGAA TGTTGGGGTGGTGCTACATTCGACGTTGCAATGAGATTCTTGCATGAGGATCCATGGGAA CGTCTGAGAAAATTAAGATCTCTGGTGCCTAATATTCCATTCCAAATGTTATTACGTGGT GCCAACGGTGTGGCTTACTCTTCATTACCTGACAATGCTATTGACCATTTTGTCAAGCAA GCCAAGGATAATGGTGTTGATATATTTAGAGTTTTTGATGCCTTGAATGATTTAGAACAA TTAAAAGTTGGTGTGAATGCTGTCAAGAAGGCCGGTGGTGTTGTCGAAGCTACTGTTTGT TACTCTGGTGACATGCTTCAGCCAGGTAAGAAATACAACTTAGACTACTACCTAGAAGTT GTTGAAAAAATAGTTCAAATGGGTACACATATCTTGGGTATTAAGGATATGGCAGGTACT ATGAAACCGGCCGCTGCCAAATTATTAATTGGCTCCCTAAGAACCAGATATCCGGATTTA CCAATTCATGTTCACAGTCATGACTCCGCAGGTACTGCTGTTGCGTCTATGACTGCATGT GCCCTAGCAGGTGCTGATGTTGTCGATGTAGCTATCAATTCAATGTCGGGCTTAACTTCC CAACCATCAATTAATGCACTGTTGGCTTCATTAGAAGGTAACATTGATACTGGGATTAAC GTTGAGCATGTTCGTGAATTAGATGCATACTGGGCCGAAATGAGACTGTTGTATTCTTGT TTCGAGGCCGACTTGAAGGGACCAGATCCAGAAGTTTACCAACATGAAATCCCAGGTGGT CAATTGACTAACTTGTTATTCCAAGCTCAACAACTGGGTCTTGGTGAACAATGGGCTGAA ACTAAAAGAGCTTACAGAGAAGCCAATTACCTACTGGGAGATATTGTTAAAGTTACCCCA ACTTCTAAGGTTGTCGGTGATTTAGCTCAATTCATGGTTTCTAACAAACTGACTTCCGAC GATATTAGACGTTTAGCTAATTCTTTGGACTTTCCTGACTCTGTTATGGACTTTTTTGAA GGTTTAATTGGTCAACCATACGGTGGGTTCCCAGAACCATTAAGATCTGATGTATTGAGA AACAAGAGAAGAAAGTTGACGTGCCGTCCAGGTTTAGAATTAGAACCATTTGATCTCGAA AAAATTAGAGAAGACTTGCAGAACAGATTCGGTGATATTGATGAATGCGATGTTGCTTCT TACAATATGTATCCAAGGGTCTATGAAGATTTCCAAAAGATCAGAGAAACATACGGTGAT TTATCAGTTCTACCAACCAAAAATTTCCTAGCACCAGCAGAACCTGATGAAGAAATCGAA GTCACCATCGAACAAGGTAAGACTTTGATTATCAAATTGCAAGCTGTTGGTGACTTAAAT AAGAAAACTGGGCAAAGAGAAGTGTATTTTGAATTGAACGGTGAATTAAGAAAGATCAGA GTTGCAGACAAGTCACAAAACATACAATCTGTTGCTAAACCAAAGGCTGATGTCCACGAT ACTCACCAAATCGGTGCACCAATGGCTGGTGTTATCATAGAAGTTAAAGTACATAAAGGG TCTTTGGTGAAAAAGGGCGAATCGATTGCTGTTTTGAGTGCCATGAAAATGGAAATGGTT GTCTCTTCACCAGCAGATGGTCAAGTTAAAGACGTTTTCATTAAGGATGGTGAAAGTGTT GACGCATCAGATTTGTTGGTTGTCCTAGAAGAAGAAACCCTACCCCCATCCCAAAAAAAG TAA | |||||||||||||||||||||||||||
| Protein Properties | ||||||||||||||||||||||||||||
| Pfam Domain Function | ||||||||||||||||||||||||||||
| Protein Residues | 1180 | |||||||||||||||||||||||||||
| Protein Molecular Weight | 130166.0 | |||||||||||||||||||||||||||
| Protein Theoretical pI | 6.47 | |||||||||||||||||||||||||||
| Signalling Regions |
| |||||||||||||||||||||||||||
| Transmembrane Regions |
| |||||||||||||||||||||||||||
| Protein Sequence | >Pyruvate carboxylase 2 MSSSKKLAGLRDNFSLLGEKNKILVANRGEIPIRIFRSAHELSMRTIAIYSHEDRLSMHR LKADEAYVIGEEGQYTPVGAYLAMDEIIEIAKKHKVDFIHPGYGFLSENSEFADKVVKAG ITWIGPPAEVIDSVGDKVSARHLAARANVPTVPGTPGPIETVQEALDFVNEYGYPVIIKA AFGGGGRGMRVVREGDDVADAFQRATSEARTAFGNGTCFVERFLDKPKHIEVQLLADNHG NVVHLFERDCSVQRRHQKVVEVAPAKTLPREVRDAILTDAVKLAKVCGYRNAGTAEFLVD NQNRHYFIEINPRIQVEHTITEEITGIDIVSAQIQIAAGATLTQLGLLQDKITTRGFSIQ CRITTEDPSKNFQPDTGRLEVYRSAGGNGVRLDGGNAYAGATISPHYDSMLVKCSCSGST YEIVRRKMIRALIEFRIRGVKTNIPFLLTLLTNPVFIEGTYWTTFIDDTPQLFQMVSSQN RAQKLLHYLADLAVNGSSIKGQIGLPKLKSNPSVPHLHDAQGNVINVTKSAPPSGWRQVL LEKGPSEFAKQVRQFNGTLLMDTTWRDAHQSLLATRVRTHDLATIAPTTAHALAGAFALE CWGGATFDVAMRFLHEDPWERLRKLRSLVPNIPFQMLLRGANGVAYSSLPDNAIDHFVKQ AKDNGVDIFRVFDALNDLEQLKVGVNAVKKAGGVVEATVCYSGDMLQPGKKYNLDYYLEV VEKIVQMGTHILGIKDMAGTMKPAAAKLLIGSLRTRYPDLPIHVHSHDSAGTAVASMTAC ALAGADVVDVAINSMSGLTSQPSINALLASLEGNIDTGINVEHVRELDAYWAEMRLLYSC FEADLKGPDPEVYQHEIPGGQLTNLLFQAQQLGLGEQWAETKRAYREANYLLGDIVKVTP TSKVVGDLAQFMVSNKLTSDDIRRLANSLDFPDSVMDFFEGLIGQPYGGFPEPLRSDVLR NKRRKLTCRPGLELEPFDLEKIREDLQNRFGDIDECDVASYNMYPRVYEDFQKIRETYGD LSVLPTKNFLAPAEPDEEIEVTIEQGKTLIIKLQAVGDLNKKTGQREVYFELNGELRKIR VADKSQNIQSVAKPKADVHDTHQIGAPMAGVIIEVKVHKGSLVKKGESIAVLSAMKMEMV VSSPADGQVKDVFIKDGESVDASDLLVVLEEETLPPSQKK | |||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||
| External Links |
| |||||||||||||||||||||||||||
| General Reference |
| |||||||||||||||||||||||||||