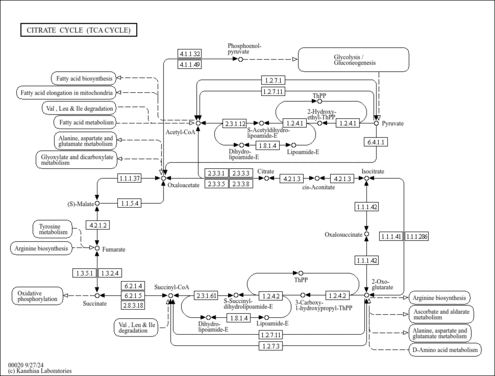
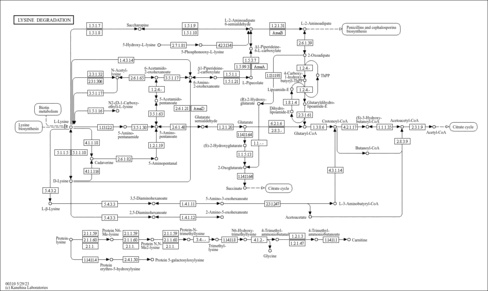
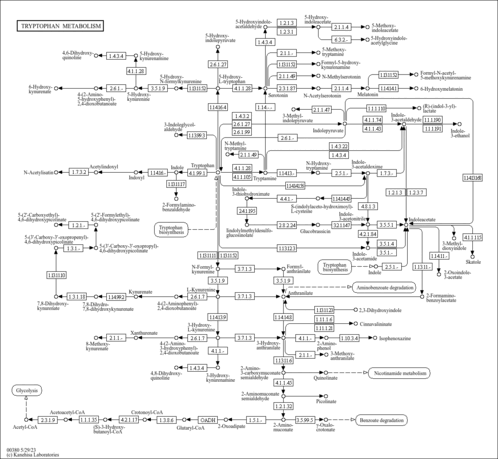
2-oxoglutarate dehydrogenase, mitochondrial (P20967)
| Identification | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | 2-oxoglutarate dehydrogenase, mitochondrial | ||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||
| Gene Name | KGD1 | ||||||||||||||||||||||||||||||
| Enzyme Class | |||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||
| General Function | Involved in oxoglutarate dehydrogenase (succinyl-transferring) activity | ||||||||||||||||||||||||||||||
| Specific Function | The 2-oxoglutarate dehydrogenase complex catalyzes the overall conversion of 2-oxoglutarate to succinyl-CoA and CO(2). It contains multiple copies of three enzymatic components:2- oxoglutarate dehydrogenase (E1), dihydrolipoamide succinyltransferase (E2) and lipoamide dehydrogenase (E3) | ||||||||||||||||||||||||||||||
| Cellular Location | Mitochondrion matrix. Mitochondrion matrix, mitochondrion nucleoid | ||||||||||||||||||||||||||||||
| SMPDB Pathways |
| ||||||||||||||||||||||||||||||
| KEGG Pathways |
| ||||||||||||||||||||||||||||||
| SMPDB Reactions |
| ||||||||||||||||||||||||||||||
| KEGG Reactions | Not Available | ||||||||||||||||||||||||||||||
| Metabolites |
| ||||||||||||||||||||||||||||||
| GO Classification |
| ||||||||||||||||||||||||||||||
| Gene Properties | |||||||||||||||||||||||||||||||
| Chromosome Location | chromosome 9 | ||||||||||||||||||||||||||||||
| Locus | YIL125W | ||||||||||||||||||||||||||||||
| Gene Sequence | >3045 bp ATGCTAAGGTTCGTGTCTTCGCAAACCTGCCGGTATAGTTCAAGAGGACTATTAAAAACA TCTTTACTTAAAAATGCATCTACTGTCAAAATTGTCGGAAGAGGGTTAGCCACCACTGGT ACAGATAATTTTCTATCGACATCAAATGCCACCTATATCGATGAAATGTACCAAGCTTGG CAAAAAGACCCATCTTCAGTCCATGTTTCATGGGACGCATATTTCAAGAATATGTCTAAC CCAAAGATTCCAGCTACAAAGGCTTTTCAGGCTCCTCCCAGTATCAGTAACTTTCCCCAG GGTACCGAAGCAGCTCCCTTAGGGACCGCAATGACTGGTTCAGTAGATGAGAACGTCTCC ATTCATCTAAAAGTGCAATTGCTATGTAGAGCTTACCAAGTTAGAGGTCATTTAAAAGCC CATATAGATCCTTTAGGGATCTCATTTGGTAGTAATAAAAATAACCCTGTTCCTCCGGAA TTGACTCTAGACTACTACGGCTTTAGCAAACACGATCTTGATAAAGAAATCAACCTAGGA CCTGGTATCCTGCCAAGGTTTGCAAGGGACGGGAAATCTAAAATGTCTCTGAAAGAGATT GTGGATCATCTAGAAAAGTTATATTGTTCCTCTTATGGGGTACAATACACACATATTCCA TCTAAGCAAAAGTGTGATTGGTTAAGAGAGAGAATTGAGATTCCTGAACCTTACCAATAT ACAGTGGACCAAAAGAGACAAATCTTAGATAGATTAACATGGGCCACTTCTTTTGAGTCA TTCTTATCTACAAAATTTCCAAATGATAAGAGGTTCGGTTTAGAAGGTTTGGAAAGTGTT GTTCCAGGTATTAAAACTTTGGTTGATCGTTCTGTTGAATTGGGTGTAGAAGATATTGTT TTGGGTATGGCTCACCGTGGTAGATTGAACGTTTTATCCAATGTGGTCCGTAAACCAAAT GAATCTATTTTTCTGAATTTAAAGGGTTCGAGCGCTCGCGATGATATTGAAGGATCGGGT GATGTCAAGTACCATTTGGGTATGAACTACCAAAGACCAACTACGTCTGGTAAGTACGTC AATTTATCGCTGGTGGCAAATCCTTCTCATTTAGAATCCCAAGATCCAGTTGTTCTTGGT AGAACTAGAGCTTTATTGCATGCCAAGAACGATTTGAAGGAAAAAACAAAGGCCTTAGGT GTGTTATTACATGGTGATGCTGCTTTTGCTGGGCAGGGTGTTGTTTATGAAACCATGGGT TTCTTGACCCTACCAGAATACTCTACTGGTGGTACTATTCATGTTATTACAAACAACCAG ATCGGATTCACTACGGATCCAAGATTTGCAAGGTCCACACCATATCCTTCCGATTTGGCT AAGGCCATTGATGCCCCAATTTTCCATGTTAACGCTAATGACGTGGAAGCTGTGACCTTT ATTTTCAATTTAGCCGCAGAATGGAGACATAAGTTCCACACAGATGCCATAATTGATGTC GTTGGTTGGAGAAAACATGGACATAATGAAACCGATCGACCATCGTTTACTCAACCATTA ATGTACAAAAAAATTGCAAAACAAAAATCTGTCATTGACGTCTATACGGAAAAATTGATA AGTGAAGGCACATTTTCTAAAAAAGATATTGATGAGCACAAGAAATGGGTATGGAACTTA TTTGAAGATGCTTTCGAAAAGACAAAGGATTACGTCCCATCTCAAAGAGAATGGTTAACT GCTGCCTGGGAAGGATTCAAATCCCCAAAGGAATTGGCCACTGAGATATTACCACATGAA CCAACTAATGTTCCAGAGAGTACTTTGAAAGAACTAGGTAAGGTACTCTCTTCGTGGCCA GAAGGTTTTGAAGTGCACAAAAATCTAAAGAGAATTTTGAAAAATAGAGGAAAATCTATT GAGACAGGTGAAGGCATCGATTGGGCCACCGGTGAAGCATTAGCGTTCGGTACATTGGTT TTGGATGGTCAGAACGTTAGGGTTTCCGGTGAAGATGTAGAAAGAGGTACATTTTCTCAA CGTCATGCAGTCTTGCATGACCAACAATCTGAAGCCATTTACACACCGCTAAGCACTCTG AATAATGAAAAGGCAGACTTCACCATTGCAAATTCCTCGTTATCTGAGTACGGTGTAATG GGTTTCGAATATGGTTATTCGCTAACCTCCCCAGATTATCTAGTCATGTGGGAGGCTCAA TTCGGTGACTTTGCAAATACAGCACAGGTTATTATTGACCAATTTATTGCCGGTGGTGAA CAAAAATGGAAGCAACGCTCTGGTTTAGTTTTGTCTTTACCCCATGGTTATGATGGCCAG GGGCCAGAACATTCGTCTGGTAGATTGGAAAGATTCTTGCAACTAGCCAATGAAGACCCA AGATATTTCCCATCTGAAGAAAAGCTACAGAGACAACATCAGGATTGTAATTTCCAGGTT GTTTATCCAACTACGCCTGCTAATTTATTCCACATTCTAAGGAGACAGCAACATCGTCAA TTCCGTAAACCATTGGCGTTATTCTTTTCTAAACAGCTGCTGCGTCACCCATTGGCCAGA TCATCTCTTTCCGAATTCACTGAAGGCGGATTCCAATGGATTATCGAAGATATTGAACAT GGAAAAAGTATTGGTACGAAAGAGGAAACCAAGAGATTAGTTTTGCTGAGTGGCCAAGTG TACACTGCCCTACATAAAAGACGTGAAAGTTTGGGTGATAAGACCACTGCTTTCTTAAAG ATTGAACAGCTGCACCCATTCCCATTTGCTCAGCTACGTGATTCATTAAATTCTTATCCA AACTTGGAAGAAATTGTTTGGTGCCAGGAAGAGCCATTGAACATGGGTTCGTGGGCATAC ACAGAACCACGCTTACACACAACATTAAAAGAAACGGATAAATATAAGGATTTCAAGGTC AGATACTGTGGTAGAAACCCAAGTGGTGCTGTTGCTGCCGGTAGCAAATCACTACATTTG GCCGAAGAAGATGCCTTTTTGAAAGATGTTTTCCAACAATCCTAA | ||||||||||||||||||||||||||||||
| Protein Properties | |||||||||||||||||||||||||||||||
| Pfam Domain Function | |||||||||||||||||||||||||||||||
| Protein Residues | 1014 | ||||||||||||||||||||||||||||||
| Protein Molecular Weight | 114416.0 | ||||||||||||||||||||||||||||||
| Protein Theoretical pI | 7.23 | ||||||||||||||||||||||||||||||
| Signalling Regions |
| ||||||||||||||||||||||||||||||
| Transmembrane Regions |
| ||||||||||||||||||||||||||||||
| Protein Sequence | >2-oxoglutarate dehydrogenase, mitochondrial MLRFVSSQTCRYSSRGLLKTSLLKNASTVKIVGRGLATTGTDNFLSTSNATYIDEMYQAW QKDPSSVHVSWDAYFKNMSNPKIPATKAFQAPPSISNFPQGTEAAPLGTAMTGSVDENVS IHLKVQLLCRAYQVRGHLKAHIDPLGISFGSNKNNPVPPELTLDYYGFSKHDLDKEINLG PGILPRFARDGKSKMSLKEIVDHLEKLYCSSYGVQYTHIPSKQKCDWLRERIEIPEPYQY TVDQKRQILDRLTWATSFESFLSTKFPNDKRFGLEGLESVVPGIKTLVDRSVELGVEDIV LGMAHRGRLNVLSNVVRKPNESIFSEFKGSSARDDIEGSGDVKYHLGMNYQRPTTSGKYV NLSLVANPSHLESQDPVVLGRTRALLHAKNDLKEKTKALGVLLHGDAAFAGQGVVYETMG FLTLPEYSTGGTIHVITNNQIGFTTDPRFARSTPYPSDLAKAIDAPIFHVNANDVEAVTF IFNLAAEWRHKFHTDAIIDVVGWRKHGHNETDQPSFTQPLMYKKIAKQKSVIDVYTEKLI SEGTFSKKDIDEHKKWVWNLFEDAFEKAKDYVPSQREWLTAAWEGFKSPKELATEILPHE PTNVPESTLKELGKVLSSWPEGFEVHKNLKRILKNRGKSIETGEGIDWATGEALAFGTLV LDGQNVRVSGEDVERGTFSQRHAVLHDQQSEAIYTPLSTLNNEKADFTIANSSLSEYGVM GFEYGYSLTSPDYLVMWEAQFGDFANTAQVIIDQFIAGGEQKWKQRSGLVLSLPHGYDGQ GPEHSSGRLERFLQLANEDPRYFPSEEKLQRQHQDCNFQVVYPTTPANLFHILRRQQHRQ FRKPLALFFSKQLLRHPLARSSLSEFTEGGFQWIIEDIEHGKSIGTKEETKRLVLLSGQV YTALHKRRESLGDKTTAFLKIEQLHPFPFAQLRDSLNSYPNLEEIVWCQEEPLNMGSWAY TEPRLHTTLKETDKYKDFKVRYCGRNPSGAVAAGSKSLHLAEEDAFLKDVFQQS | ||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||
| External Links |
| ||||||||||||||||||||||||||||||
| General Reference |
| ||||||||||||||||||||||||||||||