| Identification |
|---|
| Name | Glucosamine--fructose-6-phosphate aminotransferase [isomerizing] |
|---|
| Synonyms | - GFAT
- D-fructose-6-phosphate amidotransferase
- Hexosephosphate aminotransferase
|
|---|
| Gene Name | GFA1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in metabolic process |
|---|
| Specific Function | Involved in amino sugar synthesis (formation of chitin, supplies the amino sugars of asparagine-linked oligosaccharides of glycoproteins) |
|---|
| Cellular Location | Not Available |
|---|
| SMPDB Pathways | | Amino sugar and nucleotide sugar metabolism | PW002413 | 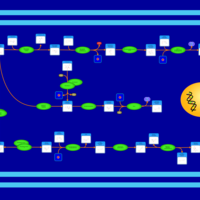 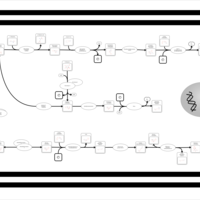 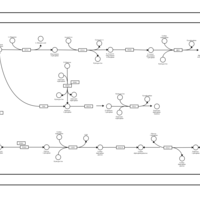 |
|
|---|
| KEGG Pathways | | Alanine, aspartate and glutamate metabolism | ec00250 |  | | Amino sugar and nucleotide sugar metabolism | ec00520 | 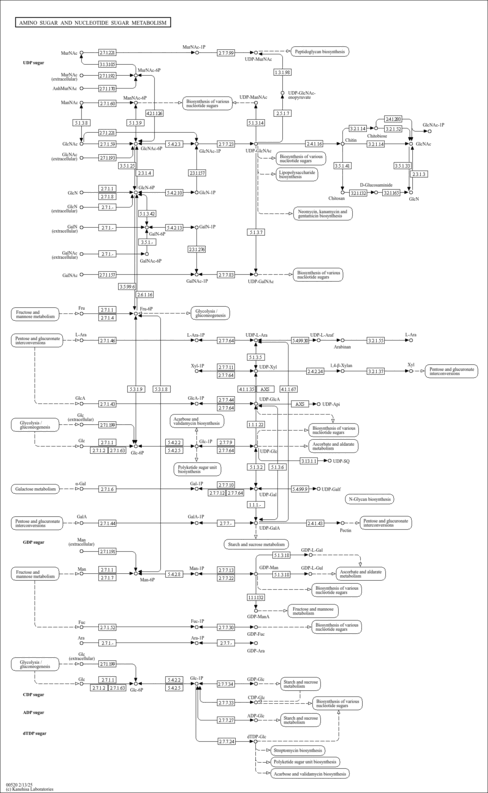 |
|
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00002 | L-Glutamine | Show | | YMDB00044 | Glucosamine 6-phosphate | Show | | YMDB00078 | Fructose 6-phosphate | Show | | YMDB00271 | L-Glutamic acid | Show | | YMDB16176 | Beta-D-Fructose 6-phosphate | Show |
|
|---|
| GO Classification | | Component |
|---|
| intracellular part | | cytoplasm | | cell part | | Function |
|---|
| catalytic activity | | transferase activity | | binding | | carbohydrate binding | | sugar binding | | transferase activity, transferring nitrogenous groups | | transaminase activity | | glutamine-fructose-6-phosphate transaminase (isomerizing) activity | | Process |
|---|
| carbohydrate biosynthetic process | | metabolic process | | primary metabolic process | | carbohydrate metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 11 |
|---|
| Locus | YKL104C |
|---|
| Gene Sequence | >2154 bp
ATGTGTGGTATCTTTGGTTACTGCAATTATCTAGTGGAAAGATCCAGAGGAGAAATTATC
GACACCTTAGTGGATGGTTTACAAAGATTAGAATATAGAGGCTATGATTCCACCGGTATT
GCTATCGATGGTGACGAAGCTGATTCTACTTTCATCTATAAGCAAATCGGTAAAGTGAGT
GCTTTGAAAGAGGAGATTACTAAGCAAAATCCGAACAGAGACGTTACTTTTGTCTCTCAT
TGTGGTATTGCGCATACTAGATGGGCTACTCACGGTCGACCAGAACAAGTTAACTGTCAC
CCTCAAAGATCTGACCCAGAAGACCAATTTGTGGTCGTTCATAATGGTATCATCACAAAT
TTTAGAGAACTGAAGACTCTTTTAATTAACAAAGGTTATAAATTCGAAAGTGATACCGAT
ACCGAGTGTATTGCTAAACTATATTTGCATTTATACAATACAAATTTACAAAATGGGCAT
GACTTAGATTTCCACGAATTAACCAAGCTAGTTCTTTTAGAACTAGAAGGTTCATACGGG
TTATTATGTAAATCTTGTCACTATCCTAATGAGGTTATCGCCACTAGAAAAGGGTCCCCT
TTACTGATTGGTGTCAAATCTGAAAAAAAACTAAAAGTCGACTTCGTGGATGTGGAATTT
CCCGAAGAAAACGCTGGTCAACCGGAAATTCCATTGAAATCTAACAACAAATCATTTGGC
TTGGGCCCAAAGAAAGCTCGTGAATTTGAAGCTGGTTCCCAAAATGCCAATTTACTACCA
ATTGCCGCCAATGAATTTAACTTGAGACATTCTCAATCCAGGGCTTTCCTATCAGAAGAT
GGATCTCCAACACCGGTGGAATTTTTTGTTTCTTCGGATGCGGCATCTGTTGTTAAACAT
ACCAAGAAGGTGCTATTTTTAGAAGATGACGATTTGGCTCATATTTACGATGGTGAGTTA
CATATTCATAGATCTAGAAGAGAAGTAGGCGCATCAATGACAAGGTCCATTCAAACTTTA
GAGATGGAGTTAGCTCAGATCATGAAGGGCCCTTACGACCATTTTATGCAAAAGGAAATC
TATGAGCAACCAGAATCTACTTTCAATACTATGAGAGGTAGAATCGACTATGAAAATAAT
AAAGTGATATTGGGTGGTTTAAAGGCATGGTTACCAGTTGTCAGAAGAGCACGGAGACTG
ATCATGATCGCATGCGGTACTTCTTATCATTCATGTTTGGCTACTCGTGCTATCTTCGAA
GAATTATCAGATATCCCAGTTAGTGTGGAATTAGCGTCTGACTTTCTGGACAGAAAATGC
CCTGTCTTCAGAGACGATGTATGCGTGTTTGTTTCACAAAGTGGTGAAACTGCGGATACC
ATGCTGGCTCTAAATTATTGTTTAGAAAGAGGAGCCTTAACTGTCGGAATTGTTAACAGT
GTTGGTTCTTCTATCTCTCGTGTCACCCACTGTGGTGTTCATATTAACGCTGGTCCTGAA
ATTGGTGTTGCCTCTACAAAAGCTTATACTTCCCAGTATATTGCCTTAGTGATGTTTGCT
CTATCGCTGTCAGATGACCGTGTATCGAAAATAGACAGAAGAATTGAAATCATTCAAGGC
TTGAAGTTAATCCCGGGCCAAATTAAGCAGGTATTAAAGCTGGAACCAAGAATAAAAAAG
CTCTGTGCGACTGAATTAAAGGATCAAAAATCTCTATTGTTATTGGGTAGAGGTTACCAA
TTTGCTGCTGCTCTGGAAGGTGCTTTGAAGATCAAAGAAATTTCTTATATGCATTCTGAA
GGTGTTTTGGCAGGTGAGTTGAAGCACGGTGTCTTGGCCTTGGTGGACGAAAACTTGCCA
ATCATTGCTTTTGGTACCAGAGACTCTCTATTCCCTAAAGTAGTTTCCTCTATTGAGCAA
GTTACTGCAAGAAAGGGCCATCCAATTATTATTTGTAACGAAAATGATGAAGTGTGGGCG
CAAAAATCTAAATCAATCGACCTGCAAACCTTAGAAGTTCCACAAACTGTTGATTGTTTA
CAAGGTCTAATTAATATTATTCCATTACAACTAATGTCATATTGGTTGGCTGTTAATAAA
GGGATTGATGTTGATTTTCCAAGAAACTTGGCTAAATCTGTTACCGTCGAATAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 717 |
|---|
| Protein Molecular Weight | 80046.0 |
|---|
| Protein Theoretical pI | 6.4 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Glucosamine--fructose-6-phosphate aminotransferase [isomerizing]
MCGIFGYCNYLVERSRGEIIDTLVDGLQRLEYRGYDSTGIAIDGDEADSTFIYKQIGKVS
ALKEEITKQNPNRDVTFVSHCGIAHTRWATHGRPEQVNCHPQRSDPEDQFVVVHNGIITN
FRELKTLLINKGYKFESDTDTECIAKLYLHLYNTNLQNGHDLDFHELTKLVLLELEGSYG
LLCKSCHYPNEVIATRKGSPLLIGVKSEKKLKVDFVDVEFPEENAGQPEIPLKSNNKSFG
LGPKKAREFEAGSQNANLLPIAANEFNLRHSQSRAFLSEDGSPTPVEFFVSSDAASVVKH
TKKVLFLEDDDLAHIYDGELHIHRSRREVGASMTRSIQTLEMELAQIMKGPYDHFMQKEI
YEQPESTFNTMRGRIDYENNKVILGGLKAWLPVVRRARRLIMIACGTSYHSCLATRAIFE
ELSDIPVSVELASDFLDRKCPVFRDDVCVFVSQSGETADTMLALNYCLERGALTVGIVNS
VGSSISRVTHCGVHINAGPEIGVASTKAYTSQYIALVMFALSLSDDRVSKIDRRIEIIQG
LKLIPGQIKQVLKLEPRIKKLCATELKDQKSLLLLGRGYQFAAALEGALKIKEISYMHSE
GVLAGELKHGVLALVDENLPIIAFGTRDSLFPKVVSSIEQVTARKGHPIIICNENDEVWA
QKSKSIDLQTLEVPQTVDCLQGLINIIPLQLMSYWLAVNKGIDVDFPRNLAKSVTVE |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Watzele, G., Tanner, W. (1989). "Cloning of the glutamine:fructose-6-phosphate amidotransferase gene from yeast. Pheromonal regulation of its transcription." J Biol Chem 264:8753-8758.2656689
- Cheret, G., Pallier, C., Valens, M., Diagnan-Fornier, B., Fukuhara, H., Bolotin-Fukuhara, M., Sor, F. (1993). "The DNA sequence analysis of the HAP4-LAP4 region on chromosome XI of Saccharomyces cerevisiae suggests the presence of a second aspartate aminotransferase gene in yeast." Yeast 9:1259-1265.8109175
- Dujon, B., Alexandraki, D., Andre, B., Ansorge, W., Baladron, V., Ballesta, J. P., Banrevi, A., Bolle, P. A., Bolotin-Fukuhara, M., Bossier, P., et, a. l. .. (1994). "Complete DNA sequence of yeast chromosome XI." Nature 369:371-378.8196765
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
- Gruhler, A., Olsen, J. V., Mohammed, S., Mortensen, P., Faergeman, N. J., Mann, M., Jensen, O. N. (2005). "Quantitative phosphoproteomics applied to the yeast pheromone signaling pathway." Mol Cell Proteomics 4:310-327.15665377
- Smolka, M. B., Albuquerque, C. P., Chen, S. H., Zhou, H. (2007). "Proteome-wide identification of in vivo targets of DNA damage checkpoint kinases." Proc Natl Acad Sci U S A 104:10364-10369.17563356
- Albuquerque, C. P., Smolka, M. B., Payne, S. H., Bafna, V., Eng, J., Zhou, H. (2008). "A multidimensional chromatography technology for in-depth phosphoproteome analysis." Mol Cell Proteomics 7:1389-1396.18407956
|
|---|

