| Identification |
|---|
| Name | Formate dehydrogenase 1 |
|---|
| Synonyms | - NAD-dependent formate dehydrogenase 1
|
|---|
| Gene Name | FDH1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
|---|
| Specific Function | Formate + NAD(+) = CO(2) + NADH |
|---|
| Cellular Location | Cytoplasm |
|---|
| SMPDB Pathways | Not Available |
|---|
| KEGG Pathways | | Glyoxylate and dicarboxylate metabolism | ec00630 | 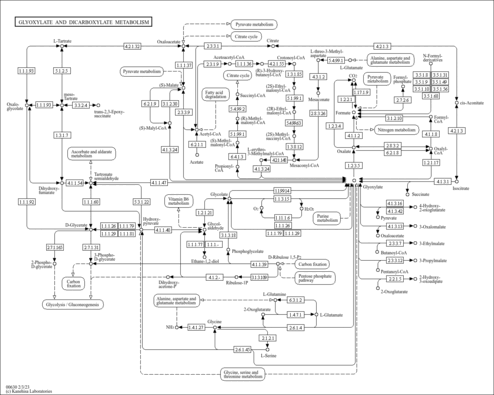 | | Methane metabolism | ec00680 | 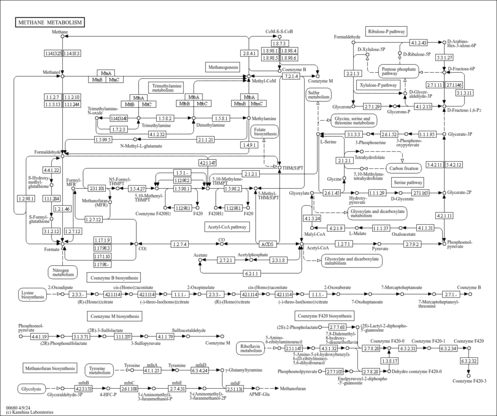 |
|
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00110 | NAD | Show | | YMDB00143 | NADH | Show | | YMDB00385 | Formic acid | Show | | YMDB00412 | NAD(+) | Show | | YMDB00912 | Carbon dioxide | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| catalytic activity | | binding | | nucleotide binding | | oxidoreductase activity | | cofactor binding | | oxidoreductase activity, acting on CH-OH group of donors | | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor | | NAD or NADH binding | | Process |
|---|
| metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 15 |
|---|
| Locus | YOR388C |
|---|
| Gene Sequence | >1131 bp
ATGTCGAAGGGAAAGGTTTTGCTGGTTCTTTACGAAGGTGGTAAGCATGCTGAAGAGCAG
GAAAAGTTATTGGGGTGTATTGAAAATGAACTTGGTATCAGAAATTTCATTGAAGAACAG
GGATACGAGTTGGTTACTACCATTGACAAGGACCCTGAGCCAACCTCAACGGTAGACAGG
GAGTTGAAAGACGCTGAAATTGTCATTACTACGCCCTTTTTCCCCGCCTACATCTCGAGA
AACAGGATTGCAGAAGCTCCTAACCTGAAGCTCTGTGTAACCGCTGGCGTCGGTTCAGAC
CATGTCGATTTAGAAGCTGCAAATGAACGGAAAATCACGGTCACCGAAGTTACTGGTTCT
AACGTCGTTTCTGTCGCAGAGCACGTTATGGCCACAATTTTGGTTTTGATAAGAAACTAT
AATGGTGGTCATCAACAAGCAATTAATGGTGAGTGGGATATTGCCGGCGTGGCTAAAAAT
GAGTATGATCTGGAAGACAAAATAATTTCAACGGTAGGTGCCGGTAGAATTGGATATAGG
GTTCTGGAAAGATTGGTCGCATTTAATCCGAAGAAGTTACTGTACTACGACTACCAGGAA
CTACCTGCGGAAGCAATCAATAGATTGAACGAGGCCAGCAAGCTTTTCAATGGCAGAGGT
GATATTGTTCAGAGAGTAGAGAAATTGGAGGATATGGTTGCTCAGTCAGATGTTGTTACC
ATCAACTGTCCATTGCACAAGGACTCAAGGGGTTTATTCAATAAAAAGCTTATTTCCCAC
ATGAAAGATGGTGCATACTTGGTGAATACCGCTAGAGGTGCTATTTGTGTCGCAGAAGAT
GTTGCCGAGGCAGTCAAGTCTGGTAAATTGGCTGGCTATGGTGGTGATGTCTGGGATAAG
CAACCAGCACCAAAAGACCATCCCTGGAGGACTATGGACAATAAGGACCACGTGGGAAAC
GCAATGACTGTTCATATCAGTGGCACATCTCTGGATGCTCAAAAGAGGTACGCTCAGGGA
GTAAAGAACATCCTAAATAGTTACTTTTCCAAAAAGTTTGATTACCGTCCACAGGATATT
ATTGTGCAGAATGGTTCTTATGCCACCAGAGCTTATGGACAGAAGAAATAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 376 |
|---|
| Protein Molecular Weight | 41714.0 |
|---|
| Protein Theoretical pI | 6.43 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Formate dehydrogenase 1
MSKGKVLLVLYEGGKHAEEQEKLLGCIENELGIRNFIEEQGYELVTTIDKDPEPTSTVDR
ELKDAEIVITTPFFPAYISRNRIAEAPNLKLCVTAGVGSDHVDLEAANERKITVTEVTGS
NVVSVAEHVMATILVLIRNYNGGHQQAINGEWDIAGVAKNEYDLEDKIISTVGAGRIGYR
VLERLVAFNPKKLLYYDYQELPAEAINRLNEASKLFNGRGDIVQRVEKLEDMVAQSDVVT
INCPLHKDSRGLFNKKLISHMKDGAYLVNTARGAICVAEDVAEAVKSGKLAGYGGDVWDK
QPAPKDHPWRTMDNKDHVGNAMTVHISGTSLDAQKRYAQGVKNILNSYFSKKFDYRPQDI
IVQNGSYATRAYGQKK |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Dujon, B., Albermann, K., Aldea, M., Alexandraki, D., Ansorge, W., Arino, J., Benes, V., Bohn, C., Bolotin-Fukuhara, M., Bordonne, R., Boyer, J., Camasses, A., Casamayor, A., Casas, C., Cheret, G., Cziepluch, C., Daignan-Fornier, B., Dang, D. V., de Haan, M., Delius, H., Durand, P., Fairhead, C., Feldmann, H., Gaillon, L., Kleine, K., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome XV." Nature 387:98-102.9169874
- van den Berg, M. A., Steensma, H. Y. (1997). "Expression cassettes for formaldehyde and fluoroacetate resistance, two dominant markers in Saccharomyces cerevisiae." Yeast 13:551-559.9178506
- Overkamp, K. M., Kotter, P., van der Hoek, R., Schoondermark-Stolk, S., Luttik, M. A., van Dijken, J. P., Pronk, J. T. (2002). "Functional analysis of structural genes for NAD(+)-dependent formate dehydrogenase in Saccharomyces cerevisiae." Yeast 19:509-520.11921099
|
|---|

