| Identification |
|---|
| Name | Aldehyde dehydrogenase [NAD(P)+] 2 |
|---|
| Synonyms | Not Available |
|---|
| Gene Name | ALD3 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in oxidoreductase activity |
|---|
| Specific Function | An aldehyde + NAD(P)(+) + H(2)O = an acid + NAD(P)H |
|---|
| Cellular Location | Not Available |
|---|
| SMPDB Pathways | |
|---|
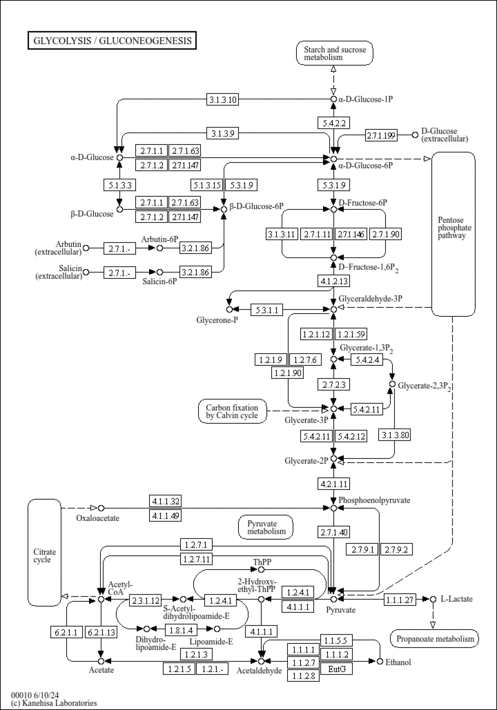
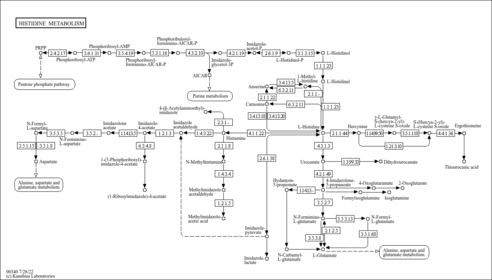
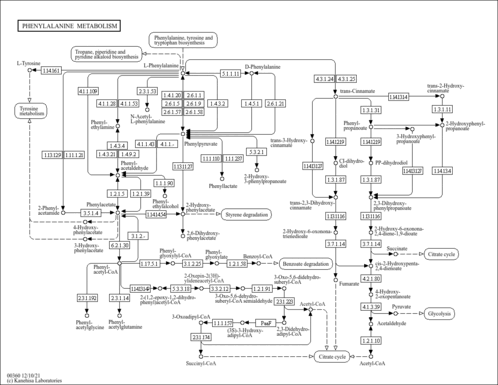
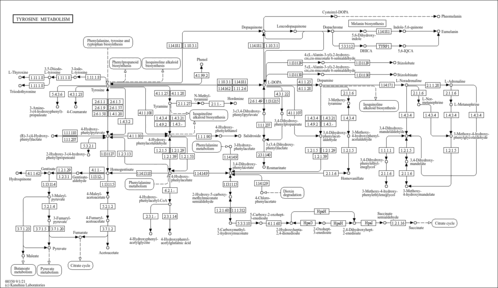
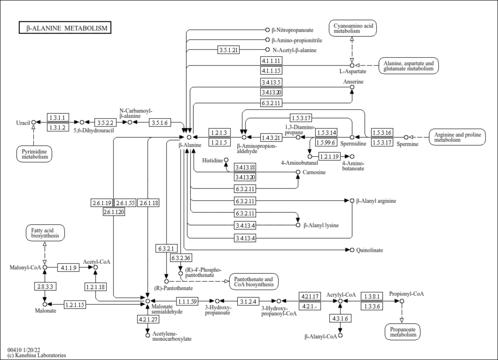
| KEGG Pathways | |
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00022 | Acetaldehyde | Show | | YMDB00048 | 3-Aminopropionaldehyde | Show | | YMDB00056 | Acetic acid | Show | | YMDB00110 | NAD | Show | | YMDB00116 | Phenylacetaldehyde | Show | | YMDB00143 | NADH | Show | | YMDB00195 | Beta-Alanine | Show | | YMDB00412 | NAD(+) | Show | | YMDB00426 | NADPH | Show | | YMDB00427 | NADP | Show | | YMDB00862 | hydron | Show | | YMDB00890 | water | Show | | YMDB00891 | Phenylacetic acid | Show | | YMDB02299 | Glyceric acid | Show | | YMDB16128 | Glyceraldehyde | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| oxidoreductase activity | | catalytic activity | | Process |
|---|
| oxidation reduction | | metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 13 |
|---|
| Locus | YMR169C |
|---|
| Gene Sequence | >1521 bp
ATGCCTACCTTGTATACTGATATCGAAATCCCACAATTGAAAATCTCTTTAAAGCAACCG
CTAGGGTTGTTTATCAACAATGAGTTTTGTCCATCATCAGATGGAAAGACCATCGAAACT
GTGAACCCAGCTACTGGCGAACCGATAACATCCTTCCAAGCAGCTAACGAAAAGGATGTA
GACAAAGCTGTGAAAGCTGCCAGGGCTGCTTTTGATAACGTTTGGTCGAAGACATCTTCT
GAGCAACGTGGTATTTATCTTTCAAACTTATTAAAACTTATTGAGGAGGAGCAAGACACA
CTTGCCGCATTAGAGACTTTAGACGCTGGTAAGCCTTTCCATTCCAATGCTAAACAAGAC
TTAGCCCAGATTATAGAACTTACAAGATACTATGCGGGGGCGGTCGACAAGTTCAATATG
GGTGAAACCATTCCATTGACTTTTAACAAGTTTGCATATACTCTAAAAGTTCCTTTTGGC
GTTGTTGCTCAAATCGTTCCATGGAATTATCCTCTAGCTATGGCTTGTAGAAAAATGCAA
GGTGCCTTAGCGGCCGGTAACACGGTTATCATCAAACCTGCTGAAAATACCTCTCTATCT
CTACTTTATTTTGCTACTTTAATTAAAAAAGCAGGTTTTCCACCTGGTGTTGTCAATGTC
ATTCCTGGTTATGGTTCCGTTGTGGGGAAAGCTTTAGGAACCCACATGGATATCGACAAA
ATATCTTTTACGGGAAGTACTAAGGTTGGCGGCTCAGTATTGGAAGCTTCCGGCCAATCG
AACCTTAAGGATATCACACTAGAATGCGGTGGTAAGTCTCCTGCTCTTGTATTTGAAGAT
GCAGACCTTGATAAGGCTATAGAATGGGTAGCAAATGGTATTTTTTTTAATTCGGGACAG
ATCTGCACTGCAAACTCAAGAGTTTATGTTCAAAGTTCGATCTACGACAAGTTTGTTGAA
AAGTTTAAAGAAACTGCAAAGAAGGAGTGGGATGTTGCAGGAAAATTTGATCCGTTTGAT
GAGAAATGCATCGTTGGTCCAGTTATATCAAGTACACAGTATGACCGCATCAAAAGTTAC
ATAGAACGTGGTAAAAAGGAGGAAAAGTTGGACATGTTCCAGACCTCTGAATTTCCTATT
GGTGGAGCTAAAGGCTACTTCATTCCCCCAACCATCTTCACTGATGTACCAGAAACATCT
AAGTTGCTGCGTGATGAAATATTTGGCCCGGTTGTGGTTGTTAGCAAGTTCACAAATTAT
GATGACGCTCTGAAGCTGGCTAATGATACTTGCTACGGGCTCGCCTCTGCGGTCTTCACC
AAAGATGTCAAGAAAGCGCACATGTTTGCTCGCGATATTAAAGCAGGAACTGTTTGGATC
AATCAAACCAATCAAGAAGAAGCTAAAGTTCCTTTTGGCGGATTTAAGATGAGTGGTATT
GGTAGAGAATCAGGCGACACCGGCGTTGATAACTATTTACAAATAAAATCAGTCCATGTG
GATCTTTCATTGGATAAATAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 506 |
|---|
| Protein Molecular Weight | 55384.80078 |
|---|
| Protein Theoretical pI | 5.42 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Aldehyde dehydrogenase [NAD(P)+] 2
MPTLYTDIEIPQLKISLKQPLGLFINNEFCPSSDGKTIETVNPATGEPITSFQAANEKDV
DKAVKAARAAFDNVWSKTSSEQRGIYLSNLLKLIEEEQDTLAALETLDAGKPFHSNAKQD
LAQIIELTRYYAGAVDKFNMGETIPLTFNKFAYTLKVPFGVVAQIVPWNYPLAMACRKMQ
GALAAGNTVIIKPAENTSLSLLYFATLIKKAGFPPGVVNVIPGYGSVVGKALGTHMDIDK
ISFTGSTKVGGSVLEASGQSNLKDITLECGGKSPALVFEDADLDKAIEWVANGIFFNSGQ
ICTANSRVYVQSSIYDKFVEKFKETAKKEWDVAGKFDPFDEKCIVGPVISSTQYDRIKSY
IERGKKEEKLDMFQTSEFPIGGAKGYFIPPTIFTDVPETSKLLRDEIFGPVVVVSKFTNY
DDALKLANDTCYGLASAVFTKDVKKAHMFARDIKAGTVWINQTNQEEAKVPFGGFKMSGI
GRESGDTGVDNYLQIKSVHVDLSLDK |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Bowman, S., Churcher, C., Badcock, K., Brown, D., Chillingworth, T., Connor, R., Dedman, K., Devlin, K., Gentles, S., Hamlin, N., Hunt, S., Jagels, K., Lye, G., Moule, S., Odell, C., Pearson, D., Rajandream, M., Rice, P., Skelton, J., Walsh, S., Whitehead, S., Barrell, B. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome XIII." Nature 387:90-93.9169872
- Navarro-Avino, J. P., Prasad, R., Miralles, V. J., Benito, R. M., Serrano, R. (1999). "A proposal for nomenclature of aldehyde dehydrogenases in Saccharomyces cerevisiae and characterization of the stress-inducible ALD2 and ALD3 genes." Yeast 15:829-842.10407263
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
|
|---|