| Identification |
|---|
| Name | Putative cystathionine beta-lyase |
|---|
| Synonyms | - CBL
- Beta-cystathionase
- Cysteine lyase
- Increased recombination centers protein 7
|
|---|
| Gene Name | IRC7 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in pyridoxal phosphate binding |
|---|
| Specific Function | L-cystathionine + H(2)O = L-homocysteine + NH(3) + pyruvate |
|---|
| Cellular Location | Not Available |
|---|
| SMPDB Pathways | |
|---|
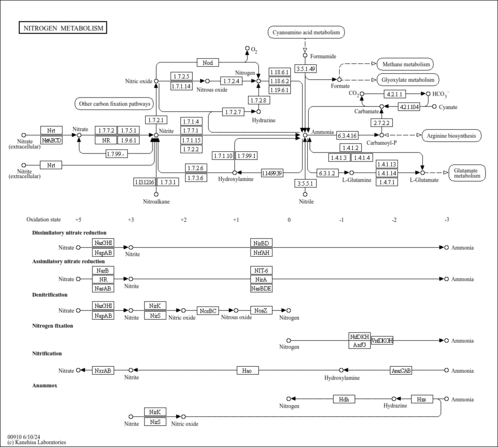
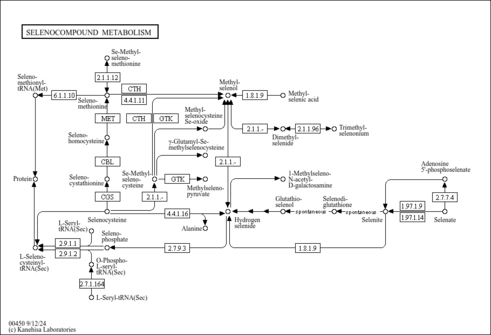
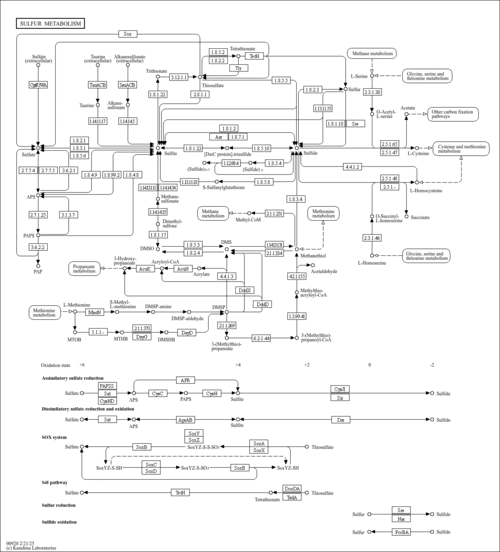
| KEGG Pathways | |
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00091 | Ammonia | Show | | YMDB00175 | Pyruvic acid | Show | | YMDB00245 | L-Cystathionine | Show | | YMDB00371 | Homocysteine | Show | | YMDB00423 | Ammonium | Show | | YMDB00520 | L-Homocysteine | Show | | YMDB00890 | water | Show | | YMDB16178 | Selenohomocysteine | Show | | YMDB16184 | Selenocystathionine | Show |
|
|---|
| GO Classification | | Component |
|---|
| intracellular part | | cytoplasm | | cell part | | Function |
|---|
| binding | | cofactor binding | | pyridoxal phosphate binding | | carbon-sulfur lyase activity | | cystathionine beta-lyase activity | | catalytic activity | | lyase activity | | Process |
|---|
| cellular metabolic process | | cellular amino acid and derivative metabolic process | | cellular amino acid metabolic process | | metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 6 |
|---|
| Locus | YFR055W |
|---|
| Gene Sequence | >1023 bp
ATGTCTACAGACAAGATCACATTTTTGTTGAACTGGCAACCAACCCCATACCATATTCCA
ATTTTCTTGGCTCAAACCAAAGGTTACTTCAAGGAGCAAGGTCTAGACATGGCCATCCTA
GAACCAACCAATCCTTCCGATGTCACTGAGTTAATTGGATCTGGTAAGGTCGACATGGGT
TTGAAAGCCATGATCCACACCTTGGCTGCCAAGGCCCGTGGTTTCCCAGTGACCTCTGTT
GCCTCTTTGTTGGACGAACCATTTACCGGTGTCTTGTACTTAAAGGGCAGTGGTATCACT
GAAGACTTCCAGTCCCTAAAGGGTAAGAAGATCGGTTACGTTGGTGAATTCGGTAAGATC
CAAATCGATGAATTGACCAAGCACTACGGTATGAAGCCAGAAGACTACACCGCCGTCAGA
TGTGGTATGAATGTCGCCAAGTACATCATCGAAGGTAAGATTGATGCCGGTATTGGTATC
GAATGTATGCAACAAGTCGAATTGGAAGAGTACTTGGCCAAGCAAGGCAGACCAGCTTCT
GATGCTAAAATGTTGAGAATTGACAAGTTGGCTTGCTTGGGTTGCTGTTGCTTCTGTACC
GTTCTTTACATCTGCAACGATGAATTTTTGAAGAAGAACCCTGAAAAGGTCAGAAAGTTC
TTGAAAGCCATCAAGAAGGCAACCGACTACGTTCTAGCCGACCCTGTGAAGGCTTGGAAA
GAATACATCGACTTCAAGCCTCAATTGAACAACGATCTATCCTACAAGCAATACCAAAGA
TGTTACGCTTACTTCTCTTCATCTTTGTACAATGTTCACCGTGACTGGAAGAAGGTTACC
GGTTACGGTAAGAGATTAGCCATCTTGCCACCAGACTATGTCTCGAACTACACTAATGAA
TACTTGTCCTGGCCAGAACCAGAAGAGGTTTCTGATCCTTTGGAAGCTCAAAGATTGATG
GCTATTCATCAAGAAAAATGCAGACAGGAAGGTACTTTCAAGAGATTGGCTCTTCCAGCT
TAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 340 |
|---|
| Protein Molecular Weight | 36971.10156 |
|---|
| Protein Theoretical pI | 6.05 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Putative cystathionine beta-lyase
MIDRTELSKFGITTQLSVIGRNPDEQSGFVNPPLYKGSTIILKKLSDLEQRKGRFYGTAG
SPTIDNLENAWTHLTGGAGTVLSASGLGSISLALLALSKAGDHILMTDSVYVPTRMLCDG
LLAKFGVETDYYDPSIGKDIEKLVKPNTTVIFLESPGSGTMEVQDIPALVSVAKKHGIKT
ILDNTWATPLFFDAHAHGIDISVEAGTKYLGGHSDLLIGLASANEECWPLLRSTYDAMAM
LPGAEDCQLALRGMRTLHLRLKEVERKALDLAAWLGNRDEVEKVLHPAFEDCPGHEYWVR
DYKGSSGLFSIVLKNGFTRAGLEKMVEGMKVLQLGFSWGG |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Murakami, Y., Naitou, M., Hagiwara, H., Shibata, T., Ozawa, M., Sasanuma, S., Sasanuma, M., Tsuchiya, Y., Soeda, E., Yokoyama, K., et, a. l. .. (1995). "Analysis of the nucleotide sequence of chromosome VI from Saccharomyces cerevisiae." Nat Genet 10:261-268.7670463
- Eki, T., Naitou, M., Hagiwara, H., Ozawa, M., Sasanuma, S. I., Sasanuma, M., Tsuchiya, Y., Shibata, T., Hanaoka, F., Murakami, Y. (1996). "Analysis of a 36.2 kb DNA sequence including the right telomere of chromosome VI from Saccharomyces cerevisiae." Yeast 12:149-167.8686379
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
|
|---|