| Identification |
|---|
| Name | Dihydroxy-acid dehydratase, mitochondrial |
|---|
| Synonyms | - DAD
- 2,3-dihydroxy acid hydrolyase
|
|---|
| Gene Name | ILV3 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in metabolic process |
|---|
| Specific Function | 2,3-dihydroxy-3-methylbutanoate = 3-methyl-2- oxobutanoate + H(2)O |
|---|
| Cellular Location | Mitochondrion |
|---|
| SMPDB Pathways | | Pantothenate and CoA biosynthesis | PW002463 |    |
|
|---|
| KEGG Pathways | | Pantothenate and CoA biosynthesis | ec00770 | 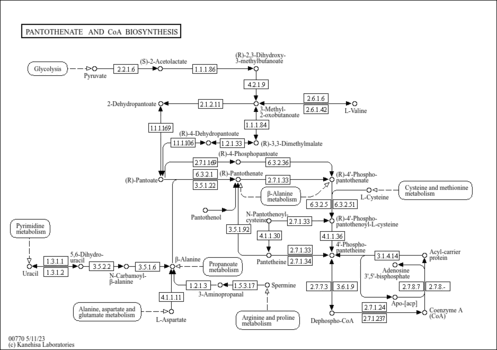 | | Valine, leucine and isoleucine biosynthesis | ec00290 | 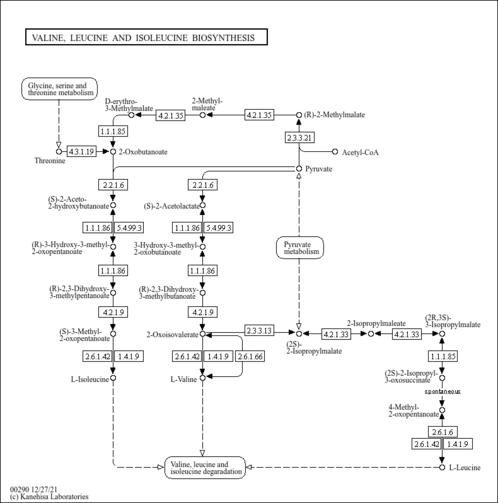 |
|
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00063 | (R)-2,3-Dihydroxy-isovalerate | Show | | YMDB00068 | (R)-2,3-Dihydroxy-3-methylvalerate | Show | | YMDB00168 | (S)-3-methyl-2-oxovaleric acid | Show | | YMDB00365 | Alpha-Ketoisovaleric acid | Show | | YMDB00890 | water | Show | | YMDB01606 | 3-Methylbutanoic acid | Show | | YMDB16286 | 2,3-Dihydroxyisovaleric acid | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| dihydroxy-acid dehydratase activity | | catalytic activity | | lyase activity | | carbon-oxygen lyase activity | | hydro-lyase activity | | Process |
|---|
| metabolic process | | cellular metabolic process | | cellular amino acid and derivative metabolic process | | cellular amino acid metabolic process | | cellular amino acid biosynthetic process | | branched chain family amino acid biosynthetic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 10 |
|---|
| Locus | YJR016C |
|---|
| Gene Sequence | >1758 bp
ATGGGCTTGTTAACGAAAGTTGCTACATCTAGACAATTCTCTACAACGAGATGCGTTGCA
AAGAAGCTCAACAAGTACTCGTATATCATCACTGAACCTAAGGGCCAAGGTGCGTCCCAG
GCCATGCTTTATGCCACCGGTTTCAAGAAGGAAGATTTCAAGAAGCCTCAAGTCGGGGTT
GGTTCCTGTTGGTGGTCCGGTAACCCATGTAACATGCATCTATTGGACTTGAATAACAGA
TGTTCTCAATCCATTGAAAAAGCGGGTTTGAAAGCTATGCAGTTCAACACCATCGGTGTT
TCAGACGGTATCTCTATGGGTACTAAAGGTATGAGATACTCGTTACAAAGTAGAGAAATC
ATTGCAGACTCCTTTGAAACCATCATGATGGCACAACACTACGATGCTAACATCGCCATC
CCATCATGTGACAAAAACATGCCCGGTGTCATGATGGCCATGGGTAGACATAACAGACCT
TCCATCATGGTATATGGTGGTACTATCTTGCCCGGTCATCCAACATGTGGTTCTTCGAAG
ATCTCTAAAAACATCGATATCGTCTCTGCGTTCCAATCCTACGGTGAATATATTTCCAAG
CAATTCACTGAAGAAGAAAGAGAAGATGTTGTGGAACATGCATGCCCAGGTCCTGGTTCT
TGTGGTGGTATGTATACTGCCAACACAATGGCTTCTGCCGCTGAAGTGCTAGGTTTGACC
ATTCCAAACTCCTCTTCCTTCCCAGCCGTTTCCAAGGAGAAGTTAGCTGAGTGTGACAAC
ATTGGTGAATACATCAAGAAGACAATGGAATTGGGTATTTTACCTCGTGATATCCTCACA
AAAGAGGCTTTTGAAAACGCCATTACTTATGTCGTTGCAACCGGTGGGTCCACTAATGCT
GTTTTGCATTTGGTGGCTGTTGCTCACTCTGCGGGTGTCAAGTTGTCACCAGATGATTTC
CAAAGAATCAGTGATACTACACCATTGATCGGTGACTTCAAACCTTCTGGTAAATACGTC
ATGGCCGATTTGATTAACGTTGGTGGTACCCAATCTGTGATTAAGTATCTATATGAAAAC
AACATGTTGCACGGTAACACAATGACTGTTACCGGTGACACTTTGGCAGAACGTGCAAAG
AAAGCACCAAGCCTACCTGAAGGACAAGAGATTATTAAGCCACTCTCCCACCCAATCAAG
GCCAACGGTCACTTGCAAATTCTGTACGGTTCATTGGCACCAGGTGGAGCTGTGGGTAAA
ATTACCGGTAAGGAAGGTACTTACTTCAAGGGTAGAGCACGTGTGTTCGAAGAGGAAGGT
GCCTTTATTGAAGCCTTGGAAAGAGGTGAAATCAAGAAGGGTGAAAAAACCGTTGTTGTT
ATCAGATATGAAGGTCCAAGAGGTGCACCAGGTATGCCTGAAATGCTAAAGCCTTCCTCT
GCTCTGATGGGTTACGGTTTGGGTAAAGATGTTGCATTGTTGACTGATGGTAGATTCTCT
GGTGGTTCTCACGGGTTCTTAATCGGCCACATTGTTCCCGAAGCCGCTGAAGGTGGTCCT
ATCGGGTTGGTCAGAGACGGCGATGAGATTATCATTGATGCTGATAATAACAAGATTGAC
CTATTAGTCTCTGATAAGGAAATGGCTCAACGTAAACAAAGTTGGGTTGCACCTCCACCT
CGTTACACAAGAGGTACTCTATCCAAGTATGCTAAGTTGGTTTCCAACGCTTCCAACGGT
TGTGTTTTAGATGCTTGA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 585 |
|---|
| Protein Molecular Weight | 62860.60156 |
|---|
| Protein Theoretical pI | 7.92 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Dihydroxy-acid dehydratase, mitochondrial
MGLLTKVATSRQFSTTRCVAKKLNKYSYIITEPKGQGASQAMLYATGFKKEDFKKPQVGV
GSCWWSGNPCNMHLLDLNNRCSQSIEKAGLKAMQFNTIGVSDGISMGTKGMRYSLQSREI
IADSFETIMMAQHYDANIAIPSCDKNMPGVMMAMGRHNRPSIMVYGGTILPGHPTCGSSK
ISKNIDIVSAFQSYGEYISKQFTEEEREDVVEHACPGPGSCGGMYTANTMASAAEVLGLT
IPNSSSFPAVSKEKLAECDNIGEYIKKTMELGILPRDILTKEAFENAITYVVATGGSTNA
VLHLVAVAHSAGVKLSPDDFQRISDTTPLIGDFKPSGKYVMADLINVGGTQSVIKYLYEN
NMLHGNTMTVTGDTLAERAKKAPSLPEGQEIIKPLSHPIKANGHLQILYGSLAPGGAVGK
ITGKEGTYFKGRARVFEEEGAFIEALERGEIKKGEKTVVVIRYEGPRGAPGMPEMLKPSS
ALMGYGLGKDVALLTDGRFSGGSHGFLIGHIVPEAAEGGPIGLVRDGDEIIIDADNNKID
LLVSDKEMAQRKQSWVAPPPRYTRGTLSKYAKLVSNASNGCVLDA |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Galibert, F., Alexandraki, D., Baur, A., Boles, E., Chalwatzis, N., Chuat, J. C., Coster, F., Cziepluch, C., De Haan, M., Domdey, H., Durand, P., Entian, K. D., Gatius, M., Goffeau, A., Grivell, L. A., Hennemann, A., Herbert, C. J., Heumann, K., Hilger, F., Hollenberg, C. P., Huang, M. E., Jacq, C., Jauniaux, J. C., Katsoulou, C., Karpfinger-Hartl, L., et, a. l. .. (1996). "Complete nucleotide sequence of Saccharomyces cerevisiae chromosome X." EMBO J 15:2031-2049.8641269
- Velasco, J. A., Cansado, J., Pena, M. C., Kawakami, T., Laborda, J., Notario, V. (1993). "Cloning of the dihydroxyacid dehydratase-encoding gene (ILV3) from Saccharomyces cerevisiae." Gene 137:179-185.8299945
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
|
|---|

