| Identification |
|---|
| Name | 6-phosphogluconate dehydrogenase, decarboxylating 1 |
|---|
| Synonyms | Not Available |
|---|
| Gene Name | GND1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in oxidoreductase activity |
|---|
| Specific Function | Catalyzes the oxidative decarboxylation of 6- phosphogluconate to ribulose 5-phosphate and CO(2), with concomitant reduction of NADP to NADPH |
|---|
| Cellular Location | Cytoplasm |
|---|
| SMPDB Pathways | |
|---|
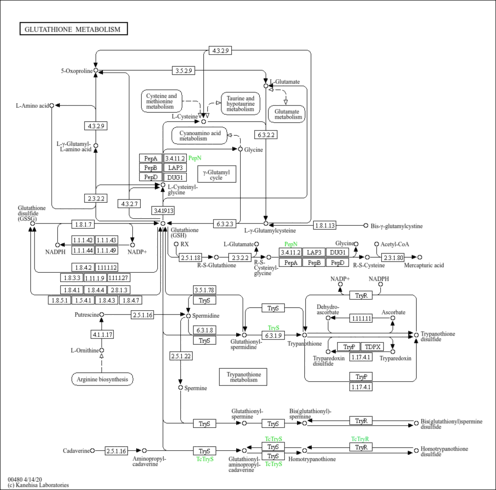
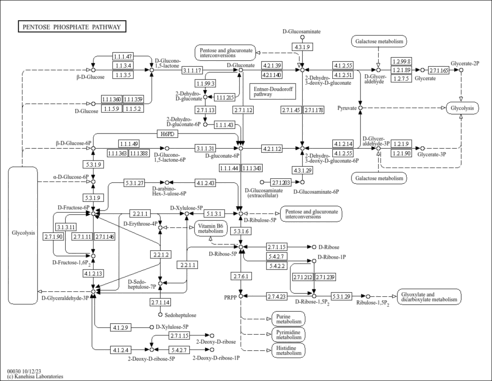
| KEGG Pathways | |
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00218 | 6-Phosphogluconic acid | Show | | YMDB00426 | NADPH | Show | | YMDB00427 | NADP | Show | | YMDB00912 | Carbon dioxide | Show | | YMDB00948 | D-ribulose 5-phosphate | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| cofactor binding | | oxidoreductase activity, acting on CH-OH group of donors | | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor | | NADP or NADPH binding | | coenzyme binding | | catalytic activity | | binding | | phosphogluconate dehydrogenase (decarboxylating) activity | | nucleotide binding | | oxidoreductase activity | | Process |
|---|
| alcohol metabolic process | | monosaccharide metabolic process | | hexose metabolic process | | glucose metabolic process | | metabolic process | | glucose catabolic process | | pentose-phosphate shunt | | oxidation reduction | | small molecule metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 8 |
|---|
| Locus | YHR183W |
|---|
| Gene Sequence | >1470 bp
ATGTCTGCTGATTTCGGTTTGATTGGTTTGGCCGTCATGGGTCAAAATTTGATCTTGAAC
GCTGCTGACCACGGTTTCACTGTTTGTGCTTACAACAGAACTCAATCCAAGGTCGACCAT
TTCTTGGCCAATGAAGCTAAGGGCAAATCTATCATCGGTGCTACTTCCATTGAAGATTTC
ATCTCCAAATTGAAGAGACCTAGAAAGGTCATGCTTTTGGTTAAAGCTGGTGCTCCAGTT
GACGCTTTGATCAACCAAATCGTCCCACTTTTGGAAAAGGGTGATATTATCATCGATGGT
GGTAACTCTCACTTCCCAGATTCTAATAGACGTTACGAAGAATTGAAGAAGAAGGGTATT
CTTTTCGTTGGTTCTGGTGTCTCCGGTGGTGAGGAAGGTGCCCGTTACGGTCCATCTTTG
ATGCCAGGTGGTTCTGAAGAAGCTTGGCCACATATTAAGAACATCTTCCAATCCATCTCT
GCTAAATCCGACGGTGAACCATGTTGCGAATGGGTTGGCCCAGCCGGTGCTGGTCACTAC
GTCAAGATGGTTCACAACGGTATTGAATACGGTGATATGCAATTGATTTGTGAAGCTTAT
GACATCATGAAGAGATTGGGTGGGTTTACCGATAAGGAAATCAGTGACGTTTTTGCCAAA
TGGAACAATGGTGTCTTGGATTCCTTCTTGGTCGAAATTACCAGAGATATTTTGAAATTC
GACGACGTCGACGGTAAGCCATTAGTTGAAAAAATCATGGATACTGCTGGTCAAAAGGGT
ACTGGTAAGTGGACTGCCATCAACGCCTTGGATTTGGGTATGCCAGTTACTTTGATTGGT
GAAGCTGTCTTTGCCCGTTGTCTATCTGCTTTGAAGAACGAGAGAATTAGAGCCTCCAAG
GTCTTACCAGGCCCAGAAGTTCCAAAAGACGCCGTCAAGGACAGAGAACAATTTGTCGAT
GATTTGGAACAAGCTTTGTATGCTTCCAAGATTATTTCTTACGCTCAAGGTTTCATGTTG
ATCCGTGAAGCTGCTGCTACTTATGGCTGGAAACTAAACAACCCTGCCATCGCTTTGATG
TGGAGAGGTGGTTGTATCATTAGATCTGTTTTCTTGGGTCAAATCACAAAGGCCTACAGA
GAAGAACCAGATTTGGAAAACTTGTTGTTCAACAAGTTCTTCGCTGATGCCGTCACCAAG
GCTCAATCTGGTTGGAGAAAGTCAATTGCGTTGGCTACCACCTACGGTATCCCAACACCA
GCCTTTTCCACCGCTTTGTCTTTCTACGATGGGTACAGATCTGAAAGATTGCCAGCCAAC
TTACTACAAGCTCAACGTGACTACTTTGGTGCTCACACTTTCAGAGTGTTGCCAGAATGT
GCTTCTGACAACTTGCCAGTAGACAAGGATATCCATATCAACTGGACTGGCCACGGTGGT
AATGTTTCTTCCTCTACATACCAAGCTTAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 489 |
|---|
| Protein Molecular Weight | 53542.69922 |
|---|
| Protein Theoretical pI | 6.6 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >6-phosphogluconate dehydrogenase, decarboxylating 1
MSADFGLIGLAVMGQNLILNAADHGFTVCAYNRTQSKVDHFLANEAKGKSIIGATSIEDF
ISKLKRPRKVMLLVKAGAPVDALINQIVPLLEKGDIIIDGGNSHFPDSNRRYEELKKKGI
LFVGSGVSGGEEGARYGPSLMPGGSEEAWPHIKNIFQSISAKSDGEPCCEWVGPAGAGHY
VKMVHNGIEYGDMQLICEAYDIMKRLGGFTDKEISDVFAKWNNGVLDSFLVEITRDILKF
DDVDGKPLVEKIMDTAGQKGTGKWTAINALDLGMPVTLIGEAVFARCLSALKNERIRASK
VLPGPEVPKDAVKDREQFVDDLEQALYASKIISYAQGFMLIREAAATYGWKLNNPAIALM
WRGGCIIRSVFLGQITKAYREEPDLENLLFNKFFADAVTKAQSGWRKSIALATTYGIPTP
AFSTALSFYDGYRSERLPANLLQAQRDYFGAHTFRVLPECASDNLPVDKDIHINWTGHGG
NVSSSTYQA |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Johnston, M., Andrews, S., Brinkman, R., Cooper, J., Ding, H., Dover, J., Du, Z., Favello, A., Fulton, L., Gattung, S., et, a. l. .. (1994). "Complete nucleotide sequence of Saccharomyces cerevisiae chromosome VIII." Science 265:2077-2082.8091229
- Garrels, J. I., Futcher, B., Kobayashi, R., Latter, G. I., Schwender, B., Volpe, T., Warner, J. R., McLaughlin, C. S. (1994). "Protein identifications for a Saccharomyces cerevisiae protein database." Electrophoresis 15:1466-1486.7895733
- Huh, W. K., Falvo, J. V., Gerke, L. C., Carroll, A. S., Howson, R. W., Weissman, J. S., O'Shea, E. K. (2003). "Global analysis of protein localization in budding yeast." Nature 425:686-691.14562095
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
- Albuquerque, C. P., Smolka, M. B., Payne, S. H., Bafna, V., Eng, J., Zhou, H. (2008). "A multidimensional chromatography technology for in-depth phosphoproteome analysis." Mol Cell Proteomics 7:1389-1396.18407956
- He, W., Wang, Y., Liu, W., Zhou, C. Z. (2007). "Crystal structure of Saccharomyces cerevisiae 6-phosphogluconate dehydrogenase Gnd1." BMC Struct Biol 7:38.17570834
|
|---|