| Identification |
|---|
| Name | Putative formate dehydrogenase 2 |
|---|
| Synonyms | - NAD-dependent formate dehydrogenase 2
|
|---|
| Gene Name | FDH2 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
|---|
| Specific Function | Formate + NAD(+) = CO(2) + NADH |
|---|
| Cellular Location | Cytoplasm |
|---|
| SMPDB Pathways | Not Available |
|---|
| KEGG Pathways | | Glyoxylate and dicarboxylate metabolism | ec00630 | 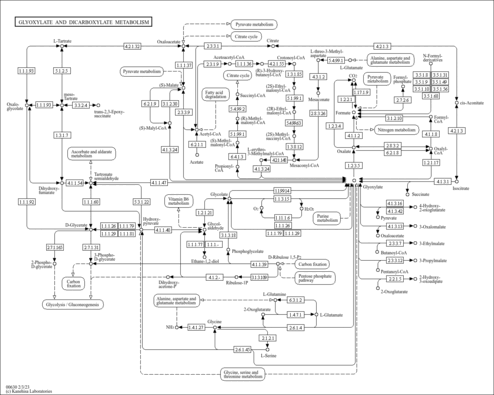 | | Methane metabolism | ec00680 | 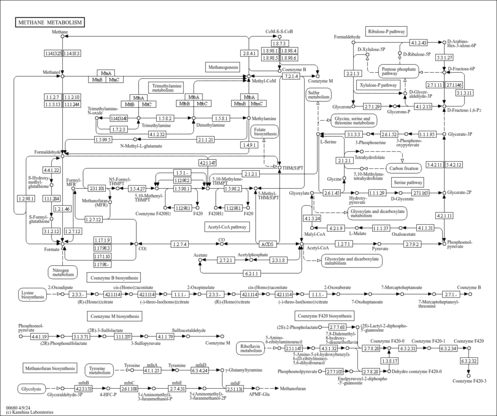 |
|
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | Not Available |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00110 | NAD | Show | | YMDB00143 | NADH | Show | | YMDB00385 | Formic acid | Show | | YMDB00412 | NAD(+) | Show | | YMDB00912 | Carbon dioxide | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| catalytic activity | | binding | | oxidoreductase activity | | cofactor binding | | oxidoreductase activity, acting on CH-OH group of donors | | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor | | Process |
|---|
| metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 16 |
|---|
| Locus | YPL275W |
|---|
| Gene Sequence | >711 bp
ATGGTGGTCATCAATAAGCAATTAATGGTGAGTGGGATATTGCCGGCGTGGCTAAAAAAT
GAGTATGATCTGGAAGACAAAATAATTTCAACGGTAGGTGCCGGTAGAATTGGATATAGG
GTTCTGGAAAGATTGGTCGCATTTAATCCGAAGAAGTTACTGTACTACGACTACCAGGAA
CTACCTGCGGAAGCAATCAATAGATTGAACGAGGCCAGCAAGCTTTTCAATGGCAGAGGT
GATATTGTTCAGAGAGTAGAGAAATTGGAGGATATGGTTGCTCAGTCAGATGTTGTTACC
ATCAACTGTCCATTGCACAAGGACTCAAGGGGTTTATTCAATAAAAAGCTTATTTCCCAC
ATGAAAGATGGTGCATACTTGGTGAATACCGCTAGAGGTGCTATTTGTGTCGCAGAAGAT
GTTGCCGAGGCAGTCAAGTCTGGTAAATTGGCTGGCTATGGTGGTGATGTCTGGGATAAG
CAACCAGCACCAAAAGACCATCCCTGGAGGACTATGGACAATAAGGACCACGTGGGAAAC
GCAATGACTGTTCATATCAGTGGCACATCTCTGCATGCTCAAAAGAGGTACGCTCAGGGA
GTAAAGAACATCCTAAATAGTTACTTTTCCAAAAAGTTTGATTACCGTCCACAGGATATT
ATTGTGCAGAATGGTTCTTATGCCACCAGAGCTTATGGACAGAAGAAATAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 236 |
|---|
| Protein Molecular Weight | 26487.19922 |
|---|
| Protein Theoretical pI | 9.71 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Putative formate dehydrogenase 2
MVVINKQLMVSGILPAWLKNEYDLEDKIISTVGAGRIGYRVLERLVAFNPKKLLYYDYQE
LPAEAINRLNEASKLFNGRGDIVQRVEKLEDMVAQSDVVTINCPLHKDSRGLFNKKLISH
MKDGAYLVNTARGAICVAEDVAEAVKSGKLAGYGGDVWDKQPAPKDHPWRTMDNKDHVGN
AMTVHISGTSLHAQKRYAQGVKNILNSYFSKKFDYRPQDIIVQNGSYATRAYGQKK |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Bussey, H., Storms, R. K., Ahmed, A., Albermann, K., Allen, E., Ansorge, W., Araujo, R., Aparicio, A., Barrell, B., Badcock, K., Benes, V., Botstein, D., Bowman, S., Bruckner, M., Carpenter, J., Cherry, J. M., Chung, E., Churcher, C., Coster, F., Davis, K., Davis, R. W., Dietrich, F. S., Delius, H., DiPaolo, T., Hani, J., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome XVI." Nature 387:103-105.9169875
- Overkamp, K. M., Kotter, P., van der Hoek, R., Schoondermark-Stolk, S., Luttik, M. A., van Dijken, J. P., Pronk, J. T. (2002). "Functional analysis of structural genes for NAD(+)-dependent formate dehydrogenase in Saccharomyces cerevisiae." Yeast 19:509-520.11921099
|
|---|

