Chitin synthase 1 (P08004)
| Identification | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Chitin synthase 1 | |||||||||||||||||
| Synonyms |
| |||||||||||||||||
| Gene Name | CHS1 | |||||||||||||||||
| Enzyme Class | ||||||||||||||||||
| Biological Properties | ||||||||||||||||||
| General Function | Involved in chitin synthase activity | |||||||||||||||||
| Specific Function | Septum formation and repair, especially under certain adverse conditions | |||||||||||||||||
| Cellular Location | Cell membrane; Multi-pass membrane protein | |||||||||||||||||
| SMPDB Pathways |
| |||||||||||||||||
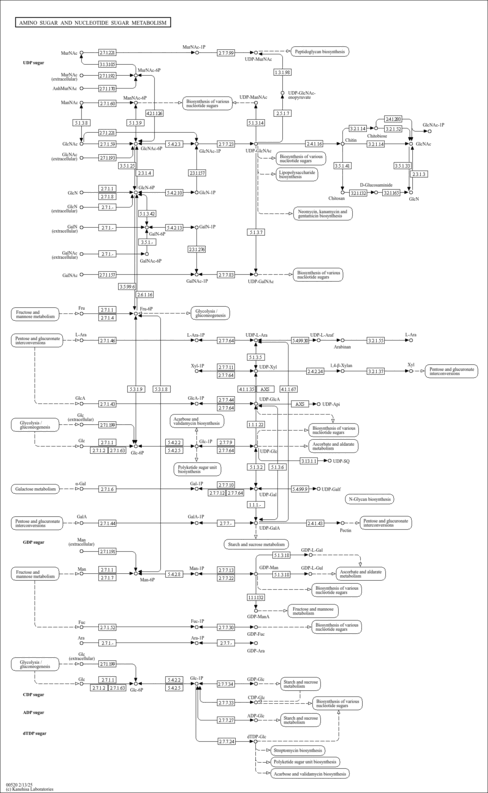
| KEGG Pathways |
| |||||||||||||||||
| SMPDB Reactions |
| |||||||||||||||||
| KEGG Reactions |
| |||||||||||||||||
| Metabolites |
| |||||||||||||||||
| GO Classification |
| |||||||||||||||||
| Gene Properties | ||||||||||||||||||
| Chromosome Location | chromosome 14 | |||||||||||||||||
| Locus | YNL192W | |||||||||||||||||
| Gene Sequence | >3396 bp ATGAGTGATCAAAATAATCGATCGAGAAATGAATATCACTCAAACCGGAAGAATGAACCT TCCTATGAACTCCAAAATGCACATAGCGGGCTATTTCACTCTTCTAATGAAGAATTAACA AACAGGAACCAAAGATATACCAATCAAAATGCCAGCATGGGTTCATTCACTCCAGTCCAA TCTTTGCAATTTCCAGAACAATCTCAGCAAACAAATATGCTTTATAACGGTGACGATGGC AATAATAATACTATCAATGATAACGAACGAGACATATATGGAGGTTTTGTCAACCACCAT CGCCAGCGTCCCCCACCAGCAACTGCAGAATACAATGACGTTTTTAATACGAATAGTCAA CAGCTACCGTCGGAACATCAATACAATAACGTACCTTCATATCCACTTCCTTCGATAAAT GTGATTCAAACCACTCCAGAACTCATACATAACGGCTCACAGACTATGGCCACCCCCATC GAAAGGCCCTTCTTTAACGAAAACGACTACTATTATAATAACAGGAACTCTAGGACGTCA CCGAGTATTGCTTCTAGTAGCGATGGTTATGCAGATCAGGAAGCTAGGCCCATTTTGGAG CAACCCAACAATAACATGAATAGCGGTAATATTCCTCAATACCATGACCAACCTTTTGGA TACAACAATGGTTACCATGGCCTACAGGCAAAAGATTACTATGACGATCCGGAGGGTGGT TATATTGATCAGAGAGGAGATGACTATCAGATTAATTCATATTTGGGTAGAAACGGTGAA ATGGTTGATCCTTACGATTATGAAAACAGTTTAAGACATATGACTCCTATGGAGCGTAGA GAATATCTTCATGATGATAGCAGACCCGTAAACGATGGAAAAGAAGAATTAGACAGTGTG AAAAGCGGTTACTCTCATAGAGACTTGGGGGAATATGACAAGGATGATTTTTCAAGGGAT GACGAGTACGATGATCTCAACACTATTGATAAATTACAGTTTCAAGCTAATGGTGTACCT GCATCATCCTCGGTGTCTTCTATCGGATCTAAAGAATCCGACATAATAGTAAGCAATGAT AACTTAACCGCAAATAGAGCACTAAAGAGAAGCGGTACTGAAATTAGGAAATTCAAACTT TGGAATGGTAATTTTGTTTTCGATTCTCCAATCAGTAAGACGCTATTGGACCAATACGCT ACTACAACAGAAAATGCAAACACTTTACCAAATGAGTTTAAGTTTATGAGATATCAAGCA GTTACTTGCGAACCTAATCAACTTGCAGAGAAGAATTTCACGGTGAGGCAGTTGAAGTAT TTAACTCCAAGGGAAACGGAATTGATGCTAGTAGTCACAATGTATAATGAAGACCATATC CTGTTAGGAAGAACTTTGAAAGGTATTATGGACAATGTCAAATATATGGTGAAAAAAAAA AATTCAAGCACTTGGGGGCCGGATGCATGGAAAAAGATTGTCGTTTGTATCATTTCAGAT GGTAGATCCAAAATTAATGAACGCTCGCTAGCATTACTAAGTTCGTTAGGTTGTTACCAG GACGGGTTTGCTAAGGATGAAATTAATGAAAAAAAAGTGGCAATGCATGTCTACGAACAT ACGACAATGATCAACATCACAAATATTTCGGAATCAGAGGTTTCATTAGAATGCAATCAA GGTACCGTTCCAATACAACTTTTGTTTTGTTTGAAAGAGCAAAATCAGAAAAAAATTAAC TCACATAGATGGGCATTTGAAGGCTTTGCAGAATTACTGCGTCCCAATATCGTTACATTG TTAGATGCTGGTACCATGCCAGGTAAAGATTCTATTTACCAGTTATGGAGAGAGTTCAGG AATCCAAATGTTGGTGGCGCATGTGGTGAAATAAGAACTGATTTGGGTAAGAGATTTGTA AAGCTTTTGAATCCTTTAGTTGCATCACAGAATTTCGAATACAAAATGTCCAATATTTTA GACAAAACAACCGAGTCTAACTTTGGATTTATTACTGTTCTACCGGGGGCATTCTCTGCG TATAGGTTTGAAGCTGTGAGAGGCCAACCATTACAGAAGTACTTTTATGGTGAAATTATG GAAAATGAAGGCTTTCATTTTTTTTCTTCCAATATGTATCTTGCTGAAGATCGTATTTTA TGCTTTGAAGTGGTCACAAAAAAAAATTGTAATTGGATTTTGAAATACTGCAGAAGTTCT TATGCTTCAACAGATGTACCGGAGAGGGTCCCTGAATTTATTCTTCAGAGGAGGCGTTGG TTGAATGGTTCATTTTTTGCTAGTGTATATTCCTTTTGTCATTTTTACAGAGTCTGGAGC AGTGGTCATAATATTGGTAGAAAACTCCTTTTGACGGTTGAATTTTTTTACCTTTTCTTC AATACATTGATTTCATGGTTTTCATTGAGTTCATTTTTCCTATTCTTTAGAATTCTCACT GTTTCTATTGCACTGGCATACCATTCAGCATTTAATGTGTTGTCCGTCATATTCCTGTGG CTTTATGGGATTTGTACCTTATCAACATTCATACTGTCATTGGGTAATAAACCTAAAAGT ACTGAGAAATTTTATGTTCTAACTTGCGTCATTTTTGCGGTGATGATGATTTACATGATA TTCTGCAGTATATTCATGAGTGTCAAATCCTTCCAAAATATATTGAAAAACGATACCATC AGCTTTGAGGGTTTGATTACCACAGAAGCTTTCAGGGATATTGTTATCTCTCTGGGCTCC ACTTATTGTTTGTACCTAATCAGTTCAATTATCTATTTGCAGCCATGGCATATGTTGACA AGTTTTATTCAGTATATTTTATTGAGTCCTTCTTACATCAATGTTTTGAATATCTATGCA TTTTGTAATGTCCACGACTTATCATGGGGTACAAAGGGTGCAATGGCAAATCCGCTGGGT AAGATTAATACTACAGAAGATGGTACGTTCAAAATGGAAGTTCTGGTCTCTAGTTCAGAG ATTCAAGCAAACTACGATAAATATTTGAAAGTTTTAAATGACTTCGATCCAAAATCAGAA TCTCGGCCTACTGAGCCATCTTATGATGAAAAAAAGACTGGCTATTATGCAAACGTTAGA TCTCTCGTGATTATCTTTTGGGTCATCACAAATTTCATCATCGTTGCTGTTGTCTTAGAA ACCGGTGGGATTGCAGATTATATTGCTATGAAATCCATATCAACTGATGACACTTTAGAA ACTGCAAAGAAGGCGGAAATTCCCTTAATGACCAGTAAGGCCTCAATTTATTTTAATGTA ATTTTATGGTTAGTTGCATTATCGGCATTAATAAGGTTCATTGGTTGCTCAATATACATG ATAGTAAGGTTTTTTAAAAAGGTTACATTTCGCTAA | |||||||||||||||||
| Protein Properties | ||||||||||||||||||
| Pfam Domain Function | ||||||||||||||||||
| Protein Residues | 1131 | |||||||||||||||||
| Protein Molecular Weight | 129918.0 | |||||||||||||||||
| Protein Theoretical pI | 6.15 | |||||||||||||||||
| Signalling Regions |
| |||||||||||||||||
| Transmembrane Regions |
| |||||||||||||||||
| Protein Sequence | >Chitin synthase 1 MSDQNNRSRNEYHSNRKNEPSYELQNAHSGLFHSSNEELTNRNQRYTNQNASMGSFTPVQ SLQFPEQSQQTNMLYNGDDGNNNTINDNERDIYGGFVNHHRQRPPPATAEYNDVFNTNSQ QLPSEHQYNNVPSYPLPSINVIQTTPELIHNGSQTMATPIERPFFNENDYYYNNRNSRTS PSIASSSDGYADQEARPILEQPNNNMNSGNIPQYHDQPFGYNNGYHGLQAKDYYDDPEGG YIDQRGDDYQINSYLGRNGEMVDPYDYENSLRHMTPMERREYLHDDSRPVNDGKEELDSV KSGYSHRDLGEYDKDDFSRDDEYDDLNTIDKLQFQANGVPASSSVSSIGSKESDIIVSND NLTANRALKRSGTEIRKFKLWNGNFVFDSPISKTLLDQYATTTENANTLPNEFKFMRYQA VTCEPNQLAEKNFTVRQLKYLTPRETELMLVVTMYNEDHILLGRTLKGIMDNVKYMVKKK NSSTWGPDAWKKIVVCIISDGRSKINERSLALLSSLGCYQDGFAKDEINEKKVAMHVYEH TTMINITNISESEVSLECNQGTVPIQLLFCLKEQNQKKINSHRWAFEGFAELLRPNIVTL LDAGTMPGKDSIYQLWREFRNPNVGGACGEIRTDLGKRFVKLLNPLVASQNFEYKMSNIL DKTTESNFGFITVLPGAFSAYRFEAVRGQPLQKYFYGEIMENEGFHFFSSNMYLAEDRIL CFEVVTKKNCNWILKYCRSSYASTDVPERVPEFILQRRRWLNGSFFASVYSFCHFYRVWS SGHNIGRKLLLTVEFFYLFFNTLISWFSLSSFFLFFRILTVSIALAYHSAFNVLSVIFLW LYGICTLSTFILSLGNKPKSTEKFYVLTCVIFAVMMIYMIFCSIFMSVKSFQNILKNDTI SFEGLITTEAFRDIVISLGSTYCLYLISSIIYLQPWHMLTSFIQYILLSPSYINVLNIYA FCNVHDLSWGTKGAMANPLGKINTTEDGTFKMEVLVSSSEIQANYDKYLKVLNDFDPKSE SRPTEPSYDEKKTGYYANVRSLVIIFWVITNFIIVAVVLETGGIADYIAMKSISTDDTLE TAKKAEIPLMTSKASIYFNVILWLVALSALIRFIGCSIYMIVRFFKKVTFR | |||||||||||||||||
| References | ||||||||||||||||||
| External Links |
| |||||||||||||||||
| General Reference |
| |||||||||||||||||