1,4-alpha-glucan-branching enzyme (P32775)
| Identification | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | 1,4-alpha-glucan-branching enzyme | ||||||||||||||
| Synonyms |
| ||||||||||||||
| Gene Name | GLC3 | ||||||||||||||
| Enzyme Class | |||||||||||||||
| Biological Properties | |||||||||||||||
| General Function | Involved in catalytic activity | ||||||||||||||
| Specific Function | Transfers a segment of a (1->4)-alpha-D-glucan chain to a primary hydroxy group in a similar glucan chain | ||||||||||||||
| Cellular Location | Cytoplasmic | ||||||||||||||
| SMPDB Pathways |
| ||||||||||||||
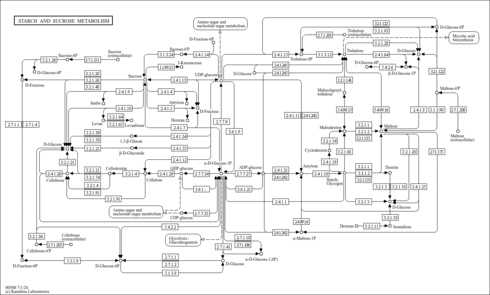
| KEGG Pathways |
| ||||||||||||||
| SMPDB Reactions |
| ||||||||||||||
| KEGG Reactions |
| ||||||||||||||
| Metabolites |
| ||||||||||||||
| GO Classification |
| ||||||||||||||
| Gene Properties | |||||||||||||||
| Chromosome Location | chromosome 5 | ||||||||||||||
| Locus | YEL011W | ||||||||||||||
| Gene Sequence | >2115 bp ATGTACAACATTCCAGATAATGTCAAAGGGGCTGTAGAATTTGATCCATGGTTGAAACCG TTTGCCGATGTACTTTCTGAAAGAAGATACCTAGCTGACAAGTGGTTGTACGACATAACT CATGCTACGCCTGATGGTTCGTATCAGTCACTCAGTAAGTTTGCTAGAGATTCATACAAG TCATATGGGTTGCATGCCAATCCGGAAACCAAAGAAATCACTTACAAAGAGTGGGCACCC AATGCGGAACGTGCATTTCTAGTGGGAGATTTCAATAACTGGGATACCACATCTCATGAG CTCAAGAACAAGGACGAATTTGGAAATTTTACAATCACACTTCATCCTTTACCTAACGGC GATTTCGCCATCCCTCATGATTCGAAGATTAAAGTCATGTTTATTTTGCCAGATGGTTCC AAGATCTTTCGTTTACCTGCGTGGATCACAAGGGCTACACAGCCTTCTAAGGAAACTTCT AAACAGTTCGGGCCCGCATATGAAGGTAGGTTTTGGAATCCAGAGAACCCCTACAAATTT GTACATCCAAGGCCAAAGTTTAGTGAATCTGTAGATTCCTTAAGGATTTATGAAGCCCAC GTCGGTATTTCGAGTCCAGAACCAAAGATAACCACATATAAAGAATTTACTGAGAAGGTC TTACCCAGAATCAAATATCTTGGCTACGATGCAATTCAATTAATGGCTATTATGGAACAT GCTTATTATGCATCATTTGGCTACCAAGTGACCAATTTTTTTGCGGCGAGCTCCCGTTTC GGCACCCCTGAGGAATTAAAGGAGCTGATTGATACTGCACATTCTATGGGCATCCTAGTT TTATTGGATGTAGTTCATAGCCATGCTTCTAAAAACGTCGAGGATGGGTTGAACATGTTT GACGGTTCTGATCATCAGTATTTCCATTCTATAAGCTCCGGTAGGGGTGAGCATCCGCTG TGGGATTCCCGTCTTTTTAATTACGGAAAATTTGAAGTGCAGCGATTCTTATTAGCAAAC CTGGCATTTTATGTTGACGTTTATCAGTTTGATGGGTTTAGATTCGATGGTGTCACATCA ATGTTATACGTTCATCATGGTGTTGGTGCCGGTGGGTCTTTTAGCGGTGACTATAACGAA TATCTTTCTCGTGACAGGTCCTTCGTCGATCATGAAGCCTTAGCTTATTTAATGTTAGCC AATGATTTGGTTCACGAAATGTTGCCAAATCTGGCTGTAACTGTTGCAGAAGATGTCTCT GGTTATCCAACTTTATGTTTACCCCGCTCCATAGGTGGTACTGGTTTCGACTACAGATTA GCAATGGCTTTGCCTGATATGTGGATCAAACTAATCAAGGAGAAGAAAGATGATGAATGG GAGATGGGAAGTATTGTATACACTTTAACTAATAGACGATATGGTGAAAAAGTGGTTGCT TATTGTGAGTCGCACGACCAAGCTTTAGTTGGTGATAAGACATTAGCGTTTTGGTTGATG GATGCCGCTATGTACACCGATATGACTGTATTGAAGGAACCATCCATCGTTATTGACCGT GGTATTGCTCTACACAAGATGATTAGGCTGATCACTCACTCGCTGGGAGGCGAGGCTTAT TTGAACTTCGAAGGTAATGAGTTCGGTCATCCAGAATGGTTGGATTTCCCTAATGTTAAC AATGGCGATACGTACAAGTATGCTCGTAGGCAGTTTAATTTGGCTGATGATCCCTTACTG CGTTACCAGAATTTAAATGAGTTTGATAGATCAATGCAATTATGTGAGAAAAGGCATAAG TGGCTGAACACGAAACAGGCTTATGTTTCATTAAAACATGAAGGTGACAAAATGATTGTG TTTGAAAGAAATAATCTGTTGTTTATTTTCAACTTCCATCCTACAAACAGCTACAGCGAT TATAGAGTTGGTGTCGAAAAAGCTGGTACTTATCACATCGTGTTGAACTCAGATCGTGCT GAATTTGGTGGACATAACAGAATCAACGAAAGTTCCGAATTCTTTACCACCGATTTGGAA TGGAACAACAGGAAGAACTTCCTCCAAGTTTATATTCCTAGTAGAGTCGCTTTGGTGCTT GCCTTAAAAGAATGA | ||||||||||||||
| Protein Properties | |||||||||||||||
| Pfam Domain Function | |||||||||||||||
| Protein Residues | 704 | ||||||||||||||
| Protein Molecular Weight | 81114.89844 | ||||||||||||||
| Protein Theoretical pI | 6.1 | ||||||||||||||
| Signalling Regions |
| ||||||||||||||
| Transmembrane Regions |
| ||||||||||||||
| Protein Sequence | >1,4-alpha-glucan-branching enzyme MYNIPDNVKGAVEFDPWLKPFADVLSERRYLADKWLYDITHATPDGSYQSLSKFARDSYK SYGLHANPETKEITYKEWAPNAERAFLVGDFNNWDTTSHELKNKDEFGNFTITLHPLPNG DFAIPHDSKIKVMFILPDGSKIFRLPAWITRATQPSKETSKQFGPAYEGRFWNPENPYKF VHPRPKFSESVDSLRIYEAHVGISSPEPKITTYKEFTEKVLPRIKYLGYDAIQLMAIMEH AYYASFGYQVTNFFAASSRFGTPEELKELIDTAHSMGILVLLDVVHSHASKNVEDGLNMF DGSDHQYFHSISSGRGEHPLWDSRLFNYGKFEVQRFLLANLAFYVDVYQFDGFRFDGVTS MLYVHHGVGAGGSFSGDYNEYLSRDRSFVDHEALAYLMLANDLVHEMLPNLAVTVAEDVS GYPTLCLPRSIGGTGFDYRLAMALPDMWIKLIKEKKDDEWEMGSIVYTLTNRRYGEKVVA YCESHDQALVGDKTLAFWLMDAAMYTDMTVLKEPSIVIDRGIALHKMIRLITHSLGGEAY LNFEGNEFGHPEWLDFPNVNNGDSYKYARRQFNLADDPLLRYQNLNEFDRSMQLCEKRHK WLNTKQAYVSLKHEGDKMIVFERNNLLFIFNFHPTNSYSDYRVGVEKAGTYHIVLNSDRA EFGGHNRINESSEFFTTDLEWNNRKNFLQVYIPSRVALVLALKE | ||||||||||||||
| References | |||||||||||||||
| External Links |
| ||||||||||||||
| General Reference |
| ||||||||||||||