Mannose-6-phosphate isomerase (P29952)
| Identification | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Mannose-6-phosphate isomerase | ||||||||||||||||||
| Synonyms |
| ||||||||||||||||||
| Gene Name | PMI40 | ||||||||||||||||||
| Enzyme Class | |||||||||||||||||||
| Biological Properties | |||||||||||||||||||
| General Function | Involved in mannose-6-phosphate isomerase activity | ||||||||||||||||||
| Specific Function | Involved in the synthesis of the GDP-mannose and dolichol-phosphate-mannose required for a number of critical mannosyl transfer reactions | ||||||||||||||||||
| Cellular Location | Cytoplasm | ||||||||||||||||||
| SMPDB Pathways |
| ||||||||||||||||||
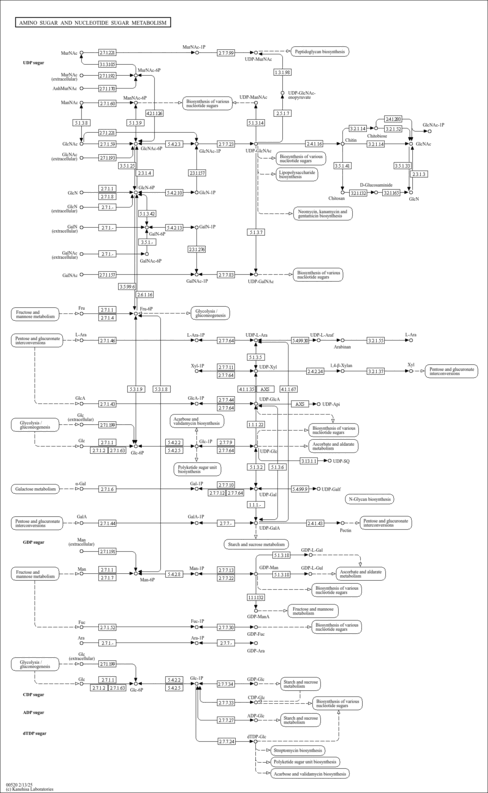
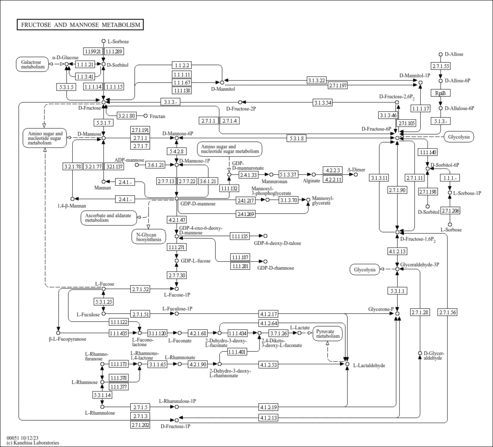
| KEGG Pathways |
| ||||||||||||||||||
| SMPDB Reactions |
| ||||||||||||||||||
| KEGG Reactions |
| ||||||||||||||||||
| Metabolites |
| ||||||||||||||||||
| GO Classification |
| ||||||||||||||||||
| Gene Properties | |||||||||||||||||||
| Chromosome Location | chromosome 5 | ||||||||||||||||||
| Locus | YER003C | ||||||||||||||||||
| Gene Sequence | >1290 bp ATGTCCAACAAGCTGTTCAGGTTAGATGCAGGCTACCAACAATACGACTGGGGTAAAATC GGCTCTTCTTCACGTGTCGCTCAATTTGCTGCCCATTCTGACCCCTCTGTTCAAATTGAA CAAGATAAACCATATGCAGAGTTATGGATGGGTACCCACAGCAAGATGCCTTCCTACAAC CATGAGTCTAAGGAATCCCTGAGAGATATCATCTCCAAGAACCCCTCTGCCATGTTAGGT AAGGACATTATTGATAAGTTCCACGCCACAAATGAATTGCCCTTCCTTTTCAAAGTTTTG TCCATTGAAAAAGTCTTGTCTATTCAAGCACATCCCGACAAAGCCTTGGGTAAAATATTG CACGCTCAAGATCCTAAGAACTATCCTGATGATAATCACAAACCTGAAATGGCCATCGCT GTGACTGACTTTGAAGGTTTCTGCGGGTTCAAACCTTTGCAAGAGATTGCAGATGAATTG AAACGTATTCCTGAATTACGCAACATTGTTGGTGAAGAAACTTCCAGGAATTTTATTGAG AACATTCAACCTTCTGCTCAGAAAGGTTCCCCAGAAGATGAGCAAAACAAAAAGCTATTG CAAGCTGTTTTCAGCAGGGTCATGAACGCTTCGGATGACAAAATCAAGATTCAAGCTCGC TCCTTGGTCGAAAGATCAAAGAATTCTCCATCAGACTTTAACAAACCTGATTTACCAGAA TTAATTCAAAGACTGAATAAACAGTTCCCTGATGACGTGGGTTTGTTTTGTGGATGTTTA TTGTTGAATCACTGCAGATTGAATGCTGGTGAAGCCATCTTTTTAAGAGCTAAGGATCCT CACGCCTATATAAGCGGTGATATTATGGAATGTATGGCTGCTTCTGACAACGTAGTTAGA GCAGGCTTCACTCCAAAATTCAAGGATGTTAAAAACTTGGTCTCCATGTTAACCTATACA TATGATCCTGTGGAAAAGCAAAAAATGCAGCCTTTAAAGTTCGACAGGTCCTCTGGTAAC GGTAAGTCAGTTTTATATAACCCTCCAATCGAAGAATTTGCTGTATTGGAGACTACTTTT GATGAGAAACTTGGTCAAAGGCATTTTGAAGGTGTTGATGGTCCAAGTATCTTAATCACT ACAAAAGGTAATGGTTACATTAAAGCAGATGGCCAAAAATTGAAAGCTGAACCCGGTTTT GTCTTTTTCATCGCTCCACATTTGCCTGTTGATTTGGAAGCTGAAGATGAGGCGTTTACT ACCTATAGAGCCTTTGTGGAACCAAATTAG | ||||||||||||||||||
| Protein Properties | |||||||||||||||||||
| Pfam Domain Function |
| ||||||||||||||||||
| Protein Residues | 429 | ||||||||||||||||||
| Protein Molecular Weight | 48188.30078 | ||||||||||||||||||
| Protein Theoretical pI | 5.83 | ||||||||||||||||||
| Signalling Regions |
| ||||||||||||||||||
| Transmembrane Regions |
| ||||||||||||||||||
| Protein Sequence | >Mannose-6-phosphate isomerase MSNKLFRLDAGYQQYDWGKIGSSSAVAQFAAHSDPSVQIEQDKPYAELWMGTHSKMPSYN HESKESLRDIISKNPSAMLGKDIIDKFHATNELPFLFKVLSIEKVLSIQAHPDKALGKIL HAQDPKNYPDDNHKPEMAIAVTDFEGFCGFKPLQEIADELKRIPELRNIVGEETSRNFIE NIQPSAQKGSPEDEQNKKLLQAVFSRVMNASDDKIKIQARSLVERSKNSPSDFNKPDLPE LIQRLNKQFPDDVGLFCGCLLLNHCRLNAGEAIFLRAKDPHAYISGDIMECMAASDNVVR AGFTPKFKDVKNLVSMLTYTYDPVEKQKMQPLKFDRSSGNGKSVLYNPPIEEFAVLETTF DEKLGQRHFEGVDGPSILITTKGNGYIKADGQKLKAEPGFVFFIAPHLPVDLEAEDEAFT TYRAFVEPN | ||||||||||||||||||
| References | |||||||||||||||||||
| External Links |
| ||||||||||||||||||
| General Reference |
| ||||||||||||||||||