| Identification |
|---|
| Name | Dihydrolipoyllysine-residue succinyltransferase component of 2-oxoglutarate dehydrogenase complex, mitochondrial |
|---|
| Synonyms | - 2-oxoglutarate dehydrogenase complex component E2
- OGDC-E2
- Dihydrolipoamide succinyltransferase component of 2-oxoglutarate dehydrogenase complex
|
|---|
| Gene Name | KGD2 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in acyltransferase activity |
|---|
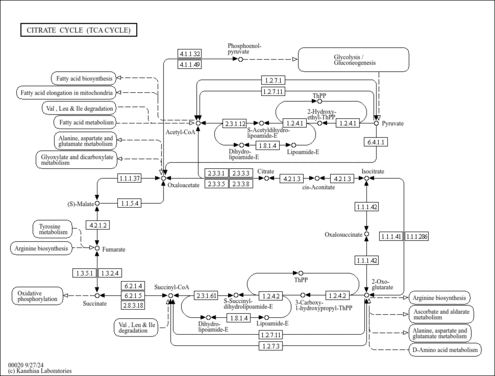
| Specific Function | The 2-oxoglutarate dehydrogenase complex catalyzes the overall conversion of 2-oxoglutarate to succinyl-CoA and CO(2). It contains multiple copies of three enzymatic components:2- oxoglutarate dehydrogenase (E1), dihydrolipoamide succinyltransferase (E2) and lipoamide dehydrogenase (E3) |
|---|
| Cellular Location | Mitochondrion |
|---|
| SMPDB Pathways | Not Available |
|---|
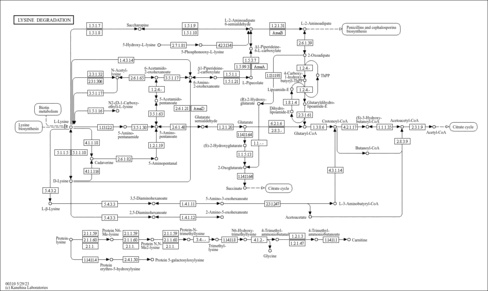
| KEGG Pathways | |
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | Not Available |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00045 | Coenzyme A | Show | | YMDB00514 | Succinyl-CoA | Show |
|
|---|
| GO Classification | | Component |
|---|
| macromolecular complex | | protein complex | | oxoglutarate dehydrogenase complex | | Function |
|---|
| S-acyltransferase activity | | S-succinyltransferase activity | | catalytic activity | | dihydrolipoyllysine-residue succinyltransferase activity | | transferase activity | | transferase activity, transferring acyl groups | | transferase activity, transferring acyl groups other than amino-acyl groups | | acyltransferase activity | | Process |
|---|
| metabolic process | | cellular metabolic process | | cofactor metabolic process | | coenzyme metabolic process | | acetyl-CoA metabolic process | | acetyl-CoA catabolic process | | tricarboxylic acid cycle |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 4 |
|---|
| Locus | YDR148C |
|---|
| Gene Sequence | >1392 bp
ATGCTTTCCAGAGCGACGCGTACTGCAGCTGCCAAATCCTTAGTAAAATCTAAAGTGGCT
AGAAATGTTATGGCTGCTTCTTTCGTCAAGAGACATGCTTCTACAAGTTTGTTCAAACAA
GCTAACAAGGTCGAATCCTTAGGTTCAATATATTTATCCGGCAAGAAAATTTCAGTTGCG
GCGAATCCGTTCTCCATAACTAGCAATCGTTTTAAATCTACCTCTATTGAAGTTCCTCCG
ATGGCAGAGTCCCTGACTGAAGGCTCTTTAAAGGAATATACTAAAAACGTTGGTGATTTT
ATTAAGGAGGACGAGCTGTTGGCCACTATTGAGACCGATAAAATTGATATTGAGGTCAAT
TCGCCAGTATCAGGTACTGTTACGAAGCTAAATTTCAAACCAGAGGACACTGTCACTGTT
GGTGAGGAGTTAGCTCAGGTCGAGCCTGGTGAAGCACCTGCTGAGGGTTCTGGAGAATCT
AAGCCAGAGCCTACCGAACAAGCGGAGCCATCGCAAGGTGTCGCCGCAAGGGAAAACTCA
AGTGAGGAAACGGCTTCAAAGAAAGAAGCTGCTCCAAAGAAAGAAGCCGCTCCAAAGAAA
GAAGTTACAGAACCAAAAAAGGCTGATCAACCAAAGAAGACCGTCTCTAAGGCGCAGGAA
CCCCCAGTAGCCTCTAACTCTTTCACACCATTTCCACGTACAGAAACCAGGGTCAAAATG
AACCGTATGAGATTGAGGATTGCCGAAAGATTAAAAGAGTCTCAAAACACTGCTGCTTCC
TTAACCACATTCAACGAAGTTGACATGTCAGCTTTGATGGAAATGAGGAAACTGTATAAA
GATGAGATTATTAAGAAGACCGGTACTAAATTCGGATTCATGGGTCTTTTCTCCAAAGCA
TGTACCTTGGCCGCCAAGGATATTCCAGCCGTCAATGGTGCCATTGAAGGTGACCAGATT
GTTTATCGTGATTACACAGATATTTCTGTTGCTGTGGCCACTCCAAAGGGTTTGGTTACC
CCCGTCGTTCGTAATGCAGAGTCATTGAGTGTTTTAGATATTGAGAACGAAATTGTTCGC
TTGAGTCATAAAGCGCGTGATGGCAAATTAACCCTAGAAGATATGACGGGTGGTACTTTC
ACCATATCTAATGGTGGTGTTTTTGGTTCATTATACGGTACTCCTATCATCAATTCACCA
CAAACAGCCGTCCTAGGCTTGCATGGTGTCAAAGAGAGACCTGTCACTGTTAATGGACAA
ATTGTCTCAAGACCAATGATGTACTTGGCTTTGACTTATGATCATAGATTGCTAGATGGT
AGAGAAGCTGTTACCTTCTTGAAGACTGTTAAAGAGTTGATTGAAGACCCTAGAAAAATG
TTGTTATGGTGA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 463 |
|---|
| Protein Molecular Weight | 50430.30078 |
|---|
| Protein Theoretical pI | 9.44 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Dihydrolipoyllysine-residue succinyltransferase component of 2-oxoglutarate dehydrogenase complex, mitochondrial
MLSRATRTAAAKSLVKSKVARNVMAASFVKRHASTSLFKQANKVESLGSIYLSGKKISVA
ANPFSITSNRFKSTSIEVPPMAESLTEGSLKEYTKNVGDFIKEDELLATIETDKIDIEVN
SPVSGTVTKLNFKPEDTVTVGEELAQVEPGEAPAEGSGESKPEPTEQAEPSQGVAARENS
SEETASKKEAAPKKEAAPKKEVTEPKKADQPKKTVSKAQEPPVASNSFTPFPRTETRVKM
NRMRLRIAERLKESQNTAASLTTFNEVDMSALMEMRKLYKDEIIKKTGTKFGFMGLFSKA
CTLAAKDIPAVNGAIEGDQIVYRDYTDISVAVATPKGLVTPVVRNAESLSVLDIENEIVR
LSHKARDGKLTLEDMTGGTFTISNGGVFGSLYGTPIINSPQTAVLGLHGVKERPVTVNGQ
IVSRPMMYLALTYDHRLLDGREAVTFLKTVKELIEDPRKMLLW |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Repetto, B., Tzagoloff, A. (1990). "Structure and regulation of KGD2, the structural gene for yeast dihydrolipoyl transsuccinylase." Mol Cell Biol 10:4221-4232.2115121
- Jacq, C., Alt-Morbe, J., Andre, B., Arnold, W., Bahr, A., Ballesta, J. P., Bargues, M., Baron, L., Becker, A., Biteau, N., Blocker, H., Blugeon, C., Boskovic, J., Brandt, P., Bruckner, M., Buitrago, M. J., Coster, F., Delaveau, T., del Rey, F., Dujon, B., Eide, L. G., Garcia-Cantalejo, J. M., Goffeau, A., Gomez-Peris, A., Zaccaria, P., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome IV." Nature 387:75-78.9169867
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
- Reinders, J., Wagner, K., Zahedi, R. P., Stojanovski, D., Eyrich, B., van der Laan, M., Rehling, P., Sickmann, A., Pfanner, N., Meisinger, C. (2007). "Profiling phosphoproteins of yeast mitochondria reveals a role of phosphorylation in assembly of the ATP synthase." Mol Cell Proteomics 6:1896-1906.17761666
|
|---|