Tryptophan synthase (P00931)
| Identification | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Tryptophan synthase | |||||||||||||||||||||||||||
| Synonyms | Not Available | |||||||||||||||||||||||||||
| Gene Name | TRP5 | |||||||||||||||||||||||||||
| Enzyme Class | ||||||||||||||||||||||||||||
| Biological Properties | ||||||||||||||||||||||||||||
| General Function | Involved in catalytic activity | |||||||||||||||||||||||||||
| Specific Function | L-serine + 1-C-(indol-3-yl)glycerol 3- phosphate = L-tryptophan + glyceraldehyde 3-phosphate + H(2)O | |||||||||||||||||||||||||||
| Cellular Location | Not Available | |||||||||||||||||||||||||||
| SMPDB Pathways |
| |||||||||||||||||||||||||||
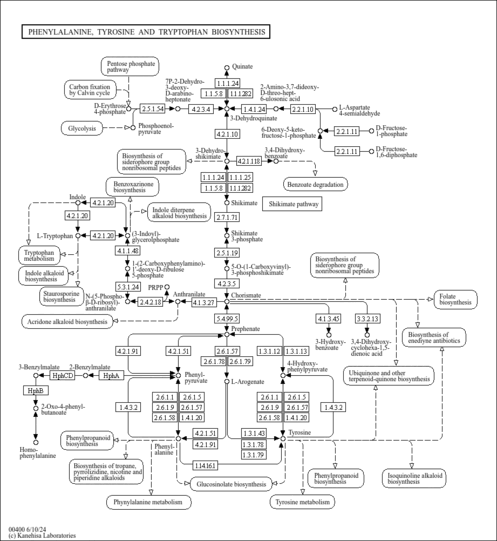
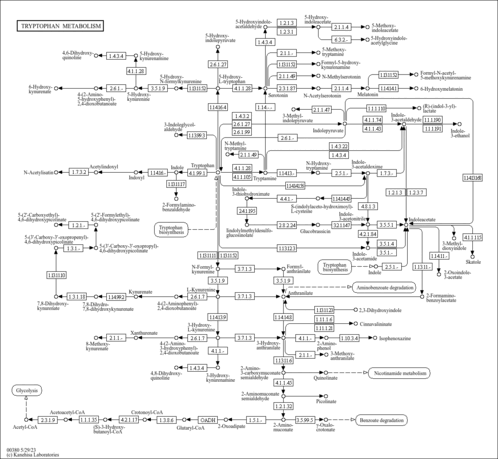
| KEGG Pathways |
| |||||||||||||||||||||||||||
| SMPDB Reactions |
| |||||||||||||||||||||||||||
| KEGG Reactions |
| |||||||||||||||||||||||||||
| Metabolites |
| |||||||||||||||||||||||||||
| GO Classification |
| |||||||||||||||||||||||||||
| Gene Properties | ||||||||||||||||||||||||||||
| Chromosome Location | chromosome 7 | |||||||||||||||||||||||||||
| Locus | YGL026C | |||||||||||||||||||||||||||
| Gene Sequence | >2124 bp ATGTCAGAACAACTCAGACAAACATTTGCTAACGCTAAAAAAGAAAACAGGAACGCCTTG GTCACATTTATGACCGCAGGTTACCCAACAGTCAAAGACACTGTCCCTATTCTCAAGGGT TTCCAGGATGGTGGTGTAGATATCATCGAATTGGGTATGCCCTTCTCTGATCCAATTGCA GATGGTCCTACAATTCAATTATCTAATACTGTGGCTTTGCAAAACGGTGTTACCTTGCCT CAAACTCTAGAAATGGTCTCCCAAGCTAGAAATGAAGGTGTTACCGTACCCATAATCCTA ATGGGTTACTATAACCCTATTCTAAACTACGGTGAAGAAAGATTTATTCAGGACGCTGCC AAGGCTGGTGCTAATGGTTTTATCATCGTCGATTTGCCACCAGAGGAGGCGTTGAAGGTC AGAAACTACATCAATGATAATGGTTTGAGCCTGATCCCACTAGTGGCTCCTTCTACCACC GACGAAAGATTGGAATTACTATCACATATTGCCGATTCGTTTGTCTACGTTGTGTCTAGA ATGGGTACTACTGGTGTTCAAAGTTCTGTGGCCAGTGATTTGGATGAACTCATCTCTAGA GTCAGAAAGTACACCAAGGATACTCCTTTGGCCGTTGGGTTTGGTGTCTCTACCAGAGAA CATTTCCAATCAGTTGGTAGTGTTGCTGACGGTGTAGTGATTGGTTCCAAAATCGTCACA TTATGTGGAGATGCTCCAGAGGGCAAAAGGTACGACGTTGCTAAGGAATATGTACAGGGA ATTCTAAATGGTGCTAAGCATAAGGTTCTGTCCAAGGACGAATTCTTTGCCTTTCAAAAA GAGTCCTTGAAGTCCGCAAACGTTAAGAAGGAAATACTGGACGAATTTGATGAAAATCAC AAGCACCCAATTAGATTTGGGGACTTTGGTGGTCAGTATGTCCCAGAAGCTCTTCATGCA TGTCTAAGAGAGTTGGAAAAGGGTTTTGATGAAGCTGTCGCCGATCCCACATTCTGGGAA GACTTCAAATCCTTGTATTCTTATATTGGCCGTCCTTCTTCACTACACAAAGCTGAGAGA TTAACTGAGCATTGTCAAGGTGCTCAAATCTGGTTGAAGAGAGAAGATCTTAACCACACG GGATCTCACAAGATCAACAATGCTTTAGCACAAGTTCTTCTAGCTAAAAGATTAGGCAAG AAGAACGTTATTGCTGAAACCGGTGCTGGTCAACACGGTGTTGCCACTGCCACTGCATGT GCTAAATTTGGCTTAACCTGTACTGTGTTCATGGGTGCAGAAGATGTTCGTCGCCAAGCT TTAAACGTCTTCAGAATGAGAATTCTCGGTGCTAAAGTAATTGCTGTTACTAATGGTACA AAGACTCTAAGAGACGCTACTTCAGAGGCATTCAGATTTTGGGTTACTAACTTGAAAACT ACTTACTACGTCGTCGGTTCTGCCATTGGTCCTCACCCATATCCAACTTTGGTTAGAACT TTCCAAAGTGTCATTGGTAAAGAAACCAAGGAACAGTTTGCTGCCATGAACAATGGTAAA TTACCTGACGCAGTTGTTGCATGTGTTGGGGGTGGTTCCAACTCTACAGGTATGTTTTCA CCATTCGAGCATGATACTTCCGTTAAGTTATTGGGTGTGGAAGCCGGTGGTGATGGTGTA GATACAAAGTTCCACTCTGCTACTCTAACTGCCGGTAGACCTGGTGTCTTCCATGGTGTC AAGACTTATGTCTTGCAAGATAGTGATGGTCAAGTCCATGATACTCATTCTGTTTCTGCT GGGTTAGACTACCCAGGTGTCGGTCCAGAATTGGCATATTGGAAATCTACTGGCCGTGCT CAATTCATTGCAGCTACTGACGCTCAGGCTCTGCTTGGCTTTAAATTATTATCTCAATTA GAAGGTATTATTCCCGCTTTGGAATCTTCTCATGCTGTTTATGGCGCTTGCGAATTGGCT AAGACGATGAAGCCTGATCAACATTTGGTTATCAATATTTCTGGTAGAGGTGATAAAGAT GTCCAAAGTGTCGCTGAAGTCTTGCCGAAATTAGGTCCAAAGATAGGTTGGGATTTGAGA TTCGAAGAAGACCCATCTGCCTAA | |||||||||||||||||||||||||||
| Protein Properties | ||||||||||||||||||||||||||||
| Pfam Domain Function | ||||||||||||||||||||||||||||
| Protein Residues | 707 | |||||||||||||||||||||||||||
| Protein Molecular Weight | 76625.5 | |||||||||||||||||||||||||||
| Protein Theoretical pI | 6.47 | |||||||||||||||||||||||||||
| Signalling Regions |
| |||||||||||||||||||||||||||
| Transmembrane Regions |
| |||||||||||||||||||||||||||
| Protein Sequence | >Tryptophan synthase MSEQLRQTFANAKKENRNALVTFMTAGYPTVKDTVPILKGFQDGGVDIIELGMPFSDPIA DGPTIQLSNTVALQNGVTLPQTLEMVSQARNEGVTVPIILMGYYNPILNYGEERFIQDAA KAGANGFIIVDLPPEEALKVRNYINDNGLSLIPLVAPSTTDERLELLSHIADSFVYVVSR MGTTGVQSSVASDLDELISRVRKYTKDTPLAVGFGVSTREHFQSVGSVADGVVIGSKIVT LCGDAPEGKRYDVAKEYVQGILNGAKHKVLSKDEFFAFQKESLKSANVKKEILDEFDENH KHPIRFGDFGGQYVPEALHACLRELEKGFDEAVADPTFWEDFKSLYSYIGRPSSLHKAER LTEHCQGAQIWLKREDLNHTGSHKINNALAQVLLAKRLGKKNVIAETGAGQHGVATATAC AKFGLTCTVFMGAEDVRRQALNVFRMRILGAKVIAVTNGTKTLRDATSEAFRFWVTNLKT TYYVVGSAIGPHPYPTLVRTFQSVIGKETKEQFAAMNNGKLPDAVVACVGGGSNSTGMFS PFEHDTSVKLLGVEAGGDGVDTKFHSATLTAGRPGVFHGVKTYVLQDSDGQVHDTHSVSA GLDYPGVGPELAYWKSTGRAQFIAATDAQALLGFKLLSQLEGIIPALESSHAVYGACELA KTMKPDQHLVINISGRGDKDVQSVAEVLPKLGPKIGWDLRFEEDPSA | |||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||
| External Links |
| |||||||||||||||||||||||||||
| General Reference |
| |||||||||||||||||||||||||||