Kynureninase (Q05979)
| Identification | |||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Kynureninase | ||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||
| Gene Name | BNA5 | ||||||||||||||||||||||||||||
| Enzyme Class | |||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||
| General Function | Involved in metabolic process | ||||||||||||||||||||||||||||
| Specific Function | Catalyzes the cleavage of L-kynurenine (L-Kyn) and L-3- hydroxykynurenine (L-3OHKyn) into anthranilic acid (AA) and 3- hydroxyanthranilic acid (3-OHAA), respectively | ||||||||||||||||||||||||||||
| Cellular Location | Cytoplasm. Nucleus | ||||||||||||||||||||||||||||
| SMPDB Pathways |
| ||||||||||||||||||||||||||||
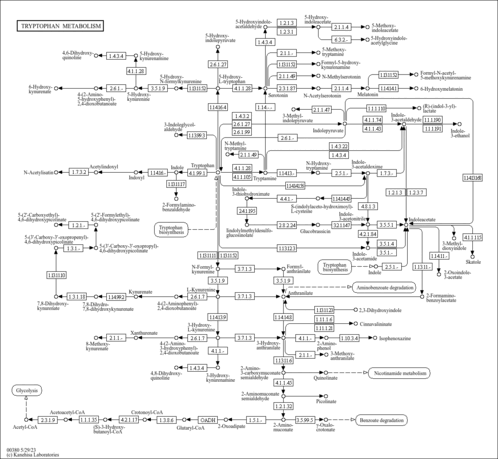
| KEGG Pathways |
| ||||||||||||||||||||||||||||
| SMPDB Reactions |
| ||||||||||||||||||||||||||||
| KEGG Reactions |
| ||||||||||||||||||||||||||||
| Metabolites |
| ||||||||||||||||||||||||||||
| GO Classification |
| ||||||||||||||||||||||||||||
| Gene Properties | |||||||||||||||||||||||||||||
| Chromosome Location | chromosome 12 | ||||||||||||||||||||||||||||
| Locus | YLR231C | ||||||||||||||||||||||||||||
| Gene Sequence | >1362 bp ATGGAGAAAGCTTTGGAATTAGACGGAGAATATCCGGAATCTCTGAGGGATGAATTCAAC ATCCCTACATTTAAATCCATGGGACTATCGTCCGACGATAAGCCTGTGACGTACTTATGC GGGAATTCTTTAGGTTTGATGCCGAAGTCAACTAGGAATTCAATTAATGCTGAGCTAGAT GCGTGGAGCGATTGTGCTGTGGAATCGCATTTCAAACATCCTGAAGAAGCCAGAGGAAAG GTGCCTTGGGTCAGCATTGACTTACCTATTCTTCCACTACTAGCCCCCATCGTGGGTGCT CAAGAAAATGAAGTTGCAGTAATGAATAGTCTCACTGCAAATTTGAATTCATTGTTAATT ACGTTTTATAAACCTACTGAGAAAAGATTCAAGATCCTTTTTGAAAAGGGCTCCTTTCCA TCAGACTATTATGCTTTCTACAACCAGTGCAAAATTCATGGAATTTCGGAACCTGAGAAT GTTTTTATTCAGATCGAGCCACGCGAGGGAGAGACTTATATCAGAACTCAAGATATCCTG GATACCATAGAGGTAAATCAAGATGAATTGGCGCTGGTCTGTTTGTCAGGTGTTCAGTAT TACACGGGGCAATATTTCGATATTGGCCGAATCACCTCATTTGCCCACCAATTCCCCGAC ATATTGGTTGGATGGGATTTAGCACACGCTGTAGGGAACGTCCCATTGCAACTTCATGAT TGGGGTGTTGACTTTGCCTGTTGGTGTTCTTACAAGTACTTGAATGCCGGGCCTGGTGGA ATTGGTGGCTTATTTGTCCACTCGAAGCATACTAAACCAGACCCTGCTAAGGAAAGTTTG CCAAGATTAGCTGGTTGGTGGGGCAATGATCCTGCTAAGCGATTTCAAATGCTGGAAGTA TTCGAGCCGATTCCAGGAGCATTGGGATTCAGGCAATCTAATCCAAGCGTTATTGACACA GTGGCATTGAGAAGTTCATTGGAATTATTTGCGAAGTTTAATGGTATTAATGAAGTTCGT AAAAGATCGTTGTTGCTGACGAACTATATGACGGAACTGTTGGAAGCCTCCAAGTATTAC AAACACCCTTTAAGGATCGAGAAATTACCATGTTTTTTTACAATATTAACTCCAACAAGC ACAGATGAAGAACATGGCGCCCAATTGTCACTTTACTTCGATTCTGACACTGGAAAAGAG GATATTATGCCCAAAGTTTTCCAATATTTGCATGACCATGGTGTCATTGGCGATGCAAGA AGACCAAACGTAATTAGATTGGCTCCTGCTCCCTTATATAATACATTTTCAGATGTATAC ATTGCGGTGAATGCACTAAATGAGGCGATGGATAAGTTGTAG | ||||||||||||||||||||||||||||
| Protein Properties | |||||||||||||||||||||||||||||
| Pfam Domain Function |
| ||||||||||||||||||||||||||||
| Protein Residues | 453 | ||||||||||||||||||||||||||||
| Protein Molecular Weight | 51031.60156 | ||||||||||||||||||||||||||||
| Protein Theoretical pI | 4.94 | ||||||||||||||||||||||||||||
| Signalling Regions |
| ||||||||||||||||||||||||||||
| Transmembrane Regions |
| ||||||||||||||||||||||||||||
| Protein Sequence | >Kynureninase MEKALELDGEYPESLRDEFNIPTFKSMGLSSDDKPVTYLCGNSLGLMPKSTRNSINAELD AWSDCAVESHFKHPEEARGKVPWVSIDLPILPLLAPIVGAQENEVAVMNSLTANLNSLLI TFYKPTEKRFKILFEKGSFPSDYYAFYNQCKIHGISEPENVFIQIEPREGETYIRTQDIL DTIEVNQDELALVCLSGVQYYTGQYFDIGRITSFAHQFPDILVGWDLAHAVGNVPLQLHD WGVDFACWCSYKYLNAGPGGIGGLFVHSKHTKPDPAKESLPRLAGWWGNDPAKRFQMLEV FEPIPGALGFRQSNPSVIDTVALRSSLELFAKFNGINEVRKRSLLLTNYMTELLEASKYY KHPLRIEKLPCFFTILTPTSTDEEHGAQLSLYFDSDTGKEDIMPKVFQYLHDHGVIGDAR RPNVIRLAPAPLYNTFSDVYIAVNALNEAMDKL | ||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||
| External Links |
| ||||||||||||||||||||||||||||
| General Reference |
| ||||||||||||||||||||||||||||