Lysophospholipase NTE1 (Q04958)
| Identification | |||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Lysophospholipase NTE1 | ||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||||||||||||||
| Gene Name | NTE1 | ||||||||||||||||||||||||||||||||||||||||||
| Enzyme Class | |||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||
| General Function | Involved in metabolic process | ||||||||||||||||||||||||||||||||||||||||||
| Specific Function | Intracellular phospholipase B that catalyzes the double deacylation of phosphatidylcholine (PC) to glycerophosphocholine (GroPCho). Plays an important role in membrane lipid homeostasis. Responsible for the rapid PC turnover in response to inositol, elevated temperatures, or when choline is present in the growth medium | ||||||||||||||||||||||||||||||||||||||||||
| Cellular Location | Endoplasmic reticulum membrane; Multi-pass membrane protein | ||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways |
| ||||||||||||||||||||||||||||||||||||||||||
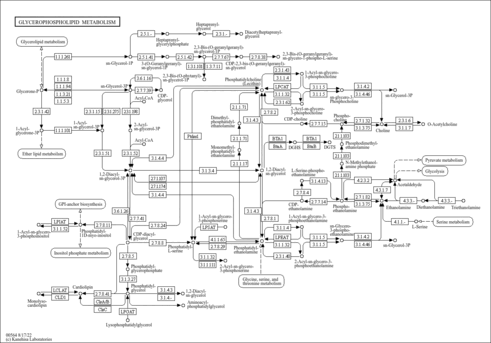
| KEGG Pathways |
| ||||||||||||||||||||||||||||||||||||||||||
| SMPDB Reactions |
| ||||||||||||||||||||||||||||||||||||||||||
| KEGG Reactions |
| ||||||||||||||||||||||||||||||||||||||||||
| Metabolites |
| ||||||||||||||||||||||||||||||||||||||||||
| GO Classification |
| ||||||||||||||||||||||||||||||||||||||||||
| Gene Properties | |||||||||||||||||||||||||||||||||||||||||||
| Chromosome Location | chromosome 13 | ||||||||||||||||||||||||||||||||||||||||||
| Locus | YML059C | ||||||||||||||||||||||||||||||||||||||||||
| Gene Sequence | >5040 bp ATGCGTTCAATGAATTGCACTACGAACAACACCAACAATACCGGCCAAAACACGAAAAAC AGTCTTGGCTCCAGCTTTAATTCCAGTAATTATACTTCTTATAGATTCCAGACATGTCTC ACCGACCAGATTATTTCTGAGGCGCAGACGTGGTCACTTTCATCCCTCTTCAACTTCTCG TGGGTTGTGTCCTACTTTGTTATGGGTGCCTCTCGTATGATCTTCAGGTACGGTTGGTAT CTGGCAACTTTATCGCTTTTAAGGATTCCGAAGTGGATCTTCTTCAAGTTACACCATGTT CAGTTCACTCTTTCCTTTTGGCTGATCCTGTTTGCCCTAGCCGTCATAGTCTTTGTGACT TACACCATCATGAAGGAGAGAATTCTATCCCAGTACAAGAGGCTGACCCCGGAGTTCTTG CCGCTGGAAAATACAGGTAAATCTGGCTCCAGCGCCAATATCAATGCAGCTTCCACTCAG TCAGCCAACGCGCCACCCGCCATAGGTTCCTCTACCACGGGTGCAAGCTCGATTATAGAC TCAAAAAAGCATTCATTGAAAGATGGCAACGAAAACGAAACCTTTCTTTCATCTTACCTT GACCAATTTCTCAGCGCCATCAAAATCTTTGGCTATCTGGAAAAACCAGTCTTCCATGAC CTGACCAAGAATATGAAAACTCAAAAAATGGATGAGGGTGAGATTTTACTGTTGGACAGT ACTATTGGGTTTGCCATCGTCGTTGAAGGCACCCTGCAGTTATATCATGAAGTAGATCAT TCAGATAAGGACCATGGCGATGAAACTGACCACAGTGACACGGATGGTCTTGACGACCAG GATAGAGATGAAGAAGACGAAGAAGAAGATGATGACATTGATAACTACGACACCAAATCT TGTTCTTCTAACCTCATCGATGAAGAAGATGAATCCGTTGGTTACATTCATCTGAAAAAT GGACTGGGAAATTTTCAACTGCTAAATACAGTCAAGCCAGGAAACCCGCTGACCTCCCTC GTGAGCATTTTAAACTTATTCACGCATTCCATGTCCTCGTATGGCAACAGTAACTTTCCC TCTGAGTTATCGTCACCTATAGACACAACGGTGTCGGTCAACAATATGTTCTGCTCATCA GAGCAGAACTTCTCCAACACAGATTCCATGACAAACTCCACCAACAGCTTTCCAACGTTC CCCTCTTCAATGCCTAAACTTGTCGCCCGTGCGGCTACAGACTGCACAATTGGTATTATC CCACCTCAATCGTTTGCCAAATTAACCGCAAAATACCCACGGTCTGCTTCCCATATCATA CAGATGGTTCTCACCAAGCTATACCATGTTACGTTCCAAACTGCTCACGACTACTTAGGT TTAACAAAGGAGATCATGGATATCGAAGTTTTACTAAATAAATCCATCGTTTATGAATTG CCTTATTATTTGAAAGAGGCCGTTATTCGTAAGTTTAAAACTGTAGATAAGTCGAGCGGA TCGGCTGATTTAGAACCGAAACCCAAAAATAGTAATGCTTCCTCTAAATTGAAGAAACCG CCCAAAGCGAAGCCATCTGATGGTATCATTCAATCGCTTAAGATTGCTAACGCCAACGCT AACACAAGTTCGAATTCCCTTTCACTGAAACCAGAATTCACCCACCATCCAAGTTCGAGA CATGTCGTACTGGGTTCAAGAGACCAATTTAATCCTGGTGATTTACTGTCAAATGTCCCA CTGTCTAGAACTATGGATATTCTGTCACCCAACCCAATCCATAATAACAACCGCAATAAA TCAAACGGTATTAACACTTCTACCTCTAACCAACATAAGCGATCTTCTAGGAGTAGCAGT AACAATGCTTCCGTACACTCTAAGAAGTTTTCTTCTTTGTCCCCAGAATTGCGCAACGCA CAACTTAGTACATCGCCCTTGTCCTTGGATAATACTTCTGTTCATGACCATATCCACCCA AGTCCCGTTCATTTGAAGGGACGGGTCTCTCCAAGACCTAATTTACTGCCCACAACATCC TTTTCTGCTGCCCAGGAAGAAACCGAGGACTCTGCTTTAAGAATGGCTTTAGTTGAGGCA ATGTTAACATACTTGGGTGTAAACAAAAGCAATATGTCCGTTTCATCTTCTTCGATAGCA AATATGTCTTCTTTAAACTCTCCTCAACTTAATGAAATGTATAGCCGCCGTCCAAGTAAC GCAAGTTTTCTGATGTCCCCTCACTGTACACCATCTGACATTTCCGTAGCTTCATCATTT GCCTCTCCTCAAACTCAACCGACAATGTTAAGAATCCTACCCAAAGAATACACTATCAGC AATAAAAGGCATAATAAAAGTAAAAGCCAGGACAAAAAAAAACCTAGAGCTTACAAAGAG GAGTTGACACCCAATTTGGATTTCGAAGATGTCAAGAAAGACTTTGCACAAGGCATTCAG TTGAAGTTCTTTAAAAAGGGCACCACAATTGTTGAGCAGAACGCTCGTGGTAAAGGCCTA TTCTATATCATTTCCGGGAAAGTTAATGTGACAACAAATTCGTCCTCGTCAGTAGTATCA TCAATGTCTAAACCAGAGCAGGTTTCAGCCCAAAGTTCACACAAGGGTGAAAATCCTCAT CATACTCAACATTTGTTATACTCTGTAGGATCTGGTGGTATTGTGGGCTACTTGTCCTCT CTAATTGGTTACAAATCATTTGTTAATATCGTTGCCAAGAGCGATGTCTATGTTGGATTT TTATCTAGTGCAACCTTAGAAAGATTGTTTGATAAATATTTCCTAATCTACTTACGGATT TCTGATTCATTAACCAAATTGCTATCTTCAAGGTTATTAAAGCTCGACCATGCTTTAGAA TGGGTTCACTTGAGGGCTTCTGAAACTCTATTTTCACAAGGTGACTCGGCTAACGGTATT TATGTTGTCCTGAATGGTAGACTAAGGCAGCTACAACAGCAGTCATTGTCAAACTCCAAT ACAAGTTCCGAAGAAGTTGAAACTCAGAACATTATTCTAGGAGAACTGGCCCAGGGGGAG AGTTTTGGTGAAGTTGAAGTGCTTACCGCTATGAATAGATATTCAACGATAGTGGCAGTG AGGGATTCTGAATTAGCGAGAATTCCGAGAACATTATTCGAACTATTAGCTCTTGAACAT CCATCTATCATGATTAGAGTTTCCAGATTAGTTGCTAAAAAAATTGTCGGAGATAGAACT GTTCCAGCATTAACAGGCGATCCATTATCTATAAAGGAAAACGACTTCACTTCTTTAATA CCTCCAACGAAAGCGTCTTACTCTAGCTCTTTAAGTCATAAACCCCAAAATATTACTTCT GGCACAATTACTTTTAGAACCATCACTATCTTACCAATTACCAGTGGACTACCTGTAGAA GCATTTGCTATGAAGTTAGTGCAAGCTTTCAAACAGGTAGGAAGGACCACGATTGGTTTA AATCAAAGAACTACCCTAACACATTTAGGTCGTCATGCATTTGACAGATTGTCCAAATTA AAACAAAGTGGTTACTTTGCTGAACTAGAAGAAATGTACCAGACTGTTGTTTATATATCA GATACACCAGTTAAATCTAATTGGACAAGAACCTGTATTGCGCAAGGTGATTGTATTCTC TTACTAGCAGATGCTAGATCTCCATCAGCCGAAATAGGAGAATATGAAAAGTTATTACTA AATTCAAAGACTACCGCAAGGACCGAGTTAATTCTTCTACACCCTGAAAGGTATGTGGAA CCAGGACTAACCCATAAGTGGTTACGCTATAGGCCATGGGTTCATTCGCATCATCACATT CAATTTAGTTTGACAGGAACAACCTTGATGAACGAAGGCAAAATGCACGTTTTGAATAAT GGTGCCTTGGCATTAATGGATAAGTTGATTCAAACAGAATTCAGTAGAAAAACTCAACAG AATATATCAAAACTACTTCCAGATTCTATCAAGAACACCGTAGAAAACTTTTCTTCAAGA TTCATGAAAAGTAAAAGACAATATTACACACCAGTTCATAGGCATAAAAATGATTTTCTA AGATTGGCTAGAATTCTTTCTGGACAAGCTATTGGACTTGTCCTCGGAGGAGGAGGTGCG AGAGGTATTAGTCATTTGGGTGTCATTCAAGCTATTGAAGAACAAGGTATTCCGGTCGAC GTTATTGGAGGAACATCGATTGGTTCCTTTGTTGGCGGTTTGTACGCGAAGGATTATGAT TTGGTGCCAATTTACGGCCGCGTGAAGAAGTTTGCTGGTCGAATTTCCTCTATATGGAGA ATGTTAACAGATTTAACCTGGCCGGTTACTTCTTATACTACTGGTCACGAATTTAATAGA GGTATTTGGAAAACTTTCGGGGATACAAGAATTGAAGACTTCTGGATCCAGTATTATTGT AATTCTACTAATATAACCGATTCGGTTCAAGAAATACATTCCTTTGGTTATGCGTGGAGG TACATCAGAGCTTCGATGTCTCTTGCTGGTCTTTTACCTCCTTTGGAAGAGAACGGGTCT ATGTTACTGGATGGTGGTTATGTTGATAATTTACCAGTAACTGAGATGAGAGCGCGTGGT TGCCAAACTATATTTGCAGTAGATGTAGGTTCTGCAGATGATAGAACGCCTATGGAATAT GGTGACTCCCTAAATGGGTTTTGGATCATTTTCAATAGATGGAATCCATTTTCTTCTCAT CCAAACATACCCAATATGGCAGAGATTCAAGTTAGATTGGGTTACGTCGCATCAGTAAAT GCTTTAGAGAAAGCGAAAAATACACCAGGTGTCGTTTATGTGAGACCACCTATTGAAGAA TATGCCACATTAGACTTTAGCAAGTTCGAAGAAATTTATCATGTGGGTGTAGACTATGGT CGTATCTTTTTGCAAGGGCTTATTGACGACGACAAGATGCCATATATACCTGGTTCACAA GAAACCACTTTGAATTCTCAAGTTCCTGAGTTTTTATTACATAGAAGAAATAGTATTTGA | ||||||||||||||||||||||||||||||||||||||||||
| Protein Properties | |||||||||||||||||||||||||||||||||||||||||||
| Pfam Domain Function | |||||||||||||||||||||||||||||||||||||||||||
| Protein Residues | 1679 | ||||||||||||||||||||||||||||||||||||||||||
| Protein Molecular Weight | 187131.0 | ||||||||||||||||||||||||||||||||||||||||||
| Protein Theoretical pI | 8.24 | ||||||||||||||||||||||||||||||||||||||||||
| Signalling Regions |
| ||||||||||||||||||||||||||||||||||||||||||
| Transmembrane Regions |
| ||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence | >Lysophospholipase NTE1 MRSMNCTTNNTNNTGQNTKNSLGSSFNSSNYTSYRFQTCLTDQIISEAQTWSLSSLFNFS WVVSYFVMGASRMIFRYGWYLATLSLLRIPKWIFFKLHHVQFTLSFWLILFALAVIVFVT YTIMKERILSQYKRLTPEFLPLENTGKSGSSANINAASTQSANAPPAIGSSTTGASSIID SKKHSLKDGNENETFLSSYLDQFLSAIKIFGYLEKPVFHDLTKNMKTQKMDEGEILLLDS TIGFAIVVEGTLQLYHEVDHSDKDHGDETDHSDTDGLDDQDRDEEDEEEDDDIDNYDTKS CSSNLIDEEDESVGYIHLKNGLGNFQLLNTVKPGNPLTSLVSILNLFTHSMSSYGNSNFP SELSSPIDTTVSVNNMFCSSEQNFSNTDSMTNSTNSFPTFPSSMPKLVARAATDCTIGII PPQSFAKLTAKYPRSASHIIQMVLTKLYHVTFQTAHDYLGLTKEIMDIEVLLNKSIVYEL PYYLKEAVIRKFKTVDKSSGSADLEPKPKNSNASSKLKKPPKAKPSDGIIQSLKIANANA NTSSNSLSLKPEFTHHPSSRHVVLGSRDQFNPGDLLSNVPLSRTMDILSPNPIHNNNRNK SNGINTSTSNQHKRSSRSSSNNASVHSKKFSSLSPELRNAQLSTSPLSLDNTSVHDHIHP SPVHLKGRVSPRPNLLPTTSFSAAQEETEDSALRMALVEAMLTYLGVNKSNMSVSSSSIA NMSSLNSPQLNEMYSRRPSNASFLMSPHCTPSDISVASSFASPQTQPTMLRILPKEYTIS NKRHNKSKSQDKKKPRAYKEELTPNLDFEDVKKDFAQGIQLKFFKKGTTIVEQNARGKGL FYIISGKVNVTTNSSSSVVSSMSKPEQVSAQSSHKGENPHHTQHLLYSVGSGGIVGYLSS LIGYKSFVNIVAKSDVYVGFLSSATLERLFDKYFLIYLRISDSLTKLLSSRLLKLDHALE WVHLRASETLFSQGDSANGIYVVLNGRLRQLQQQSLSNSNTSSEEVETQNIILGELAQGE SFGEVEVLTAMNRYSTIVAVRDSELARIPRTLFELLALEHPSIMIRVSRLVAKKIVGDRT VPALTGDPLSIKENDFTSLIPPTKASYSSSLSHKPQNITSGTITFRTITILPITSGLPVE AFAMKLVQAFKQVGRTTIGLNQRTTLTHLGRHAFDRLSKLKQSGYFAELEEMYQTVVYIS DTPVKSNWTRTCIAQGDCILLLADARSPSAEIGEYEKLLLNSKTTARTELILLHPERYVE PGLTHKWLRYRPWVHSHHHIQFSLTGTTLMNEGKMHVLNNGALALMDKLIQTEFSRKTQQ NISKLLPDSIKNTVENFSSRFMKSKRQYYTPVHRHKNDFLRLARILSGQAIGLVLGGGGA RGISHLGVIQAIEEQGIPVDVIGGTSIGSFVGGLYAKDYDLVPIYGRVKKFAGRISSIWR MLTDLTWPVTSYTTGHEFNRGIWKTFGDTRIEDFWIQYYCNSTNITDSVQEIHSFGYAWR YIRASMSLAGLLPPLEENGSMLLDGGYVDNLPVTEMRARGCQTIFAVDVGSADDRTPMEY GDSLNGFWIIFNRWNPFSSHPNIPNMAEIQVRLGYVASVNALEKAKNTPGVVYVRPPIEE YATLDFSKFEEIYHVGVDYGRIFLQGLIDDDKMPYIPGSQETTLNSQVPEFLLHRRNSI | ||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||
| External Links |
| ||||||||||||||||||||||||||||||||||||||||||
| General Reference |
| ||||||||||||||||||||||||||||||||||||||||||