Glycerophosphodiester phosphodiesterase GDE1 (Q02979)
| Identification | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Glycerophosphodiester phosphodiesterase GDE1 | |||||||||||||||||||||||||||
| Synonyms | Not Available | |||||||||||||||||||||||||||
| Gene Name | GDE1 | |||||||||||||||||||||||||||
| Enzyme Class | ||||||||||||||||||||||||||||
| Biological Properties | ||||||||||||||||||||||||||||
| General Function | Involved in protein binding | |||||||||||||||||||||||||||
| Specific Function | Glycerophosphocholine glycerophosphodiesterase responsible for the hydrolysis of intracellular glycerophosphocholine into glycerol-phosphate and choline. The choline is used for phosphatidyl-choline synthesis. Required for utilization of glycerophosphocholine as phosphate source | |||||||||||||||||||||||||||
| Cellular Location | Cytoplasm | |||||||||||||||||||||||||||
| SMPDB Pathways |
| |||||||||||||||||||||||||||
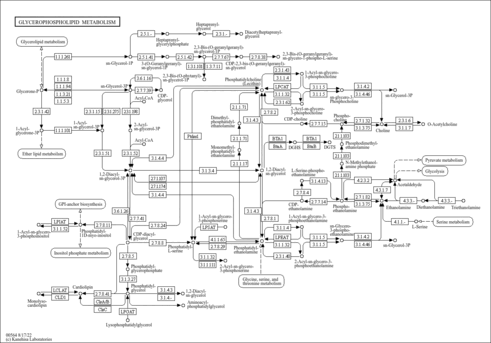
| KEGG Pathways |
| |||||||||||||||||||||||||||
| SMPDB Reactions |
| |||||||||||||||||||||||||||
| KEGG Reactions |
| |||||||||||||||||||||||||||
| Metabolites |
| |||||||||||||||||||||||||||
| GO Classification |
| |||||||||||||||||||||||||||
| Gene Properties | ||||||||||||||||||||||||||||
| Chromosome Location | chromosome 16 | |||||||||||||||||||||||||||
| Locus | YPL110C | |||||||||||||||||||||||||||
| Gene Sequence | >3672 bp ATGAAGTTCGGAAAAACCTTTGCCAATCATCGCATTCCAGAGTGGTCAAGTCAATATGTT GGCTACAAGTCGTTGAAGAAAATGATTAAAGAAATTACAAGACTTCAAGAAGACATATAT AGGGCACATAACAAGAATAGCTATGATGAAGGCAGGCCACCAACAAAAATGAGAGATTCA TCTAATAGTGCTCAAAATTATTTGGACTCTCCAAAAATCCAAAAGTTATTGGCGTCGTTT TTCTTTGCAGTTGACAGGGATATTGAGAAGGTTGATACATTTTACAATTCCCAGTATGCG GAATATAAGAAAAGATTTGAGAGATTGCTTTCTTCTAATCAATTCAATGAGATAAAATCA ACATTAGTGGTCGATGCCAACAAAGAAGATGCAGTAGCGCAAACCTTACTCACAAAGGAT ACGAGAGAGATGAATATGCTTTTAAAGGGAACAAGTCAAGCCAGCCGTCTATCTTACCAT AAAGATGACTTAATTGAAATTCAGTCTATCTTAGCTGAGTTAAGAAAACAATTTAGAAAC TTAAAGTGGTACGCTGAACTAAATAAGAGAGCTTTTGGTAAGATTTTAAAGAAACTGGAT AAAAAAGTCGGCACAAATCAGCAAATGTCAACTATGAAAACTAGAATCTTGCCGTTACAA TTTGCTAATGATTCTCTTATTACCAAAGATTTGAGTTTGCTAAAAACTATCTGGGAACAA GTGACGTTCAGGATAAATTCTTACGAGAGAGTAATGAGGAGTACCTCGCCGAACGCAAAT GCTAATGATAATACAGAATTCTTTAAGATTATTTGCGTTTTTATAGAAGAAGATGATAGC AAAGGATTGATTAGAGAGCTAACAAACCTATATTCGGAATTGTCTTTGATTCCAACAAGA ATTATGATAAGTGTTTTAAACAAGGCAGCACTGTCTAAATCACTGGCATGTATTGATGCT ATCTTAAAGGTTATTCCGTCTCTAAATGACTCGGAAGATATTAACAGAAGAAATTTTTTC CATCATCACATAATTGCAATCGGGAAGTTAATCAGAAAACAGGAAATTTTAAGTAGGAAA AAGAAGTCACAACCCTCAAAATATACTAATAGTGAGGGAGAAATTGTAACAGATTTAAGA ACCCTGCATACGACCCTATCTGCCCCGGCAGAATCAGATAGCATTACAGAGGAGGAAAAA AGTTCGGCCTGTACACTCTCTTATATTTTAGAGGAGTTACCAATTCATTTAAGACCATGC CTTTTTCAGCATGATAATTACAAGAGAACGCCCTTGCATTATTCTTGCCAGTATGGCTTA TCTGAAGTTACGAAGTTGATTATCAAGTTAATGAAAGAATGGAATATCTGGAATGAAATC CCTATTGATGACGTTTCCGCGTTCGGAGATGCCGAATCATTGACACCTTTACATTTATGT GTGTTGGGTGCTCATCCCAAAACAACAGAAGTCTTATTGCAATCTTTGGATCCGAACGTG AAATTAAAAAGTTCAAGTTTGTTACATTTGGCCACTGAGTGGAATAACTATCCCCTTTTA CATGTTCTCCTATCATCAAAAAGGTTTGATATTAATTATCAAGACAACGAATTACATGAA ACACCGCTATATCTTGCTTGTAGATTAAATTTTTTCGAAGCTGCTGTGTGTTTGTTGTAC AATGGAGCCGACCTTGAAATAAGAGAAAAACTTTTTGGCTGGACAGCTATCTTCGTTGCA GCAGCAGAAGGGTTTACGGACATTGTAAAGCTCTTGATAGCCAACAACGCGAACTTTGAT ATTGAAGATGAAGGTGGGTGGACGCCGATGGAGCACGCGGTATTGCGCGGTCATTTGCAC ATTGCTGATATGGTTCAAATCAGAGACGAACTGGTCACGCATCCCCATTCACAGTTGAAC AGTGGTAGCGAAGAGAAAGAACCACTAAACGAAATTAGTGCCGGAGAATTAAATGAGAGA AATGAAAATGGAAACGGAGGTAATAAAGGGTCCTTGGGAAAACTGGCCGGACCAATAAAG TCATATGGACACAGATTCCTGGATAATAATGAATCACTTATACTAATAACATTGGGAAGT AATGACACAAGAAACAAAAGCCCTTCAATTTCATTAAGCTCGGAAGCACTAGCAAAAGTT ATTGGGTTGGAAACAGATTGTGCATTATCGTTAGTAATTTCATGCAATGATTCAATTGAT AAATCTTCTGTAATCTTAGATTTACCATTAGATGATAATGTGGACGCAGTAGACTTCAAA GTGCCTTTCAAAGTTGATTATTCTCATACATTATATTTTGATATTGTGCCAACATATGGA ACACGCTCGTTGGAGACCCATAATAGAATAGATTGTCAAAAAAATAACAATAATTACGTA ATGGCAAGGGGTGTTTCCATGTTGAATAAGTCGTACAGCTCAGTGGGAGTTAATAGAAGT ATATTAAATGGTAGCGTGACGGTCCCCATCATCGCTAATCATACACTGGAGATACTTGGG ACGCTGAAATTTGAATATATTATTATCACACCATTTGAACATCCACAACTGCCCTTAGAA AGAACAGAAACGTACTGGAAATCCTTAGTGTCCACTCGTGTTATAGGACATAGAGGCCTG GGTAAAAATAATCCTAATAAATCATTACAAATTGGAGAGAACACAGTCGAGTCTTTCATC ATGGCAGCCTCATTGGGAGCATCGTATGTAGAGTTTGATGTTCAGTTGACGAAGGATAAC GTACCAGTGGTATATCACGATTTTCTCGTAGCTGAAACAGGTGTCGATATCCCCATGCAT GAATTGACGCTGGAACAGTTTTTAGATTTGAACAATGCCGATAAAGAGCACATTCAACGT GGAGCAGGTCACTCACCTCATCATGTAAATGGAGCAGATACAGCACTACAAAAATATAGA GGGCGTTCGGTAGATGATTCAGACGTTAGTACATTGAGGAGAGCTTGGGATTTACACGAC AACGACCCGAATGGGAAGTCCAACAATGCTCATTGGTCTGACAATCGAATGAGGCTAACC AAAACATTTAAAAAGAACAATTTTAAGGGAAACGCTAGAGGACATTCAATAGCTTCATCA TTTGTTACCTTGAAAGAACTATTCAAGAAAATCCCTGCCAACGTTGGGTTTAACATAGAA TGTAAGTTTCCCATGCTTGATGAGGCAGAAGAGGAAGAGTTAGGGCAAATAATGATGGAA ATGAACCATTGGGTTGACACCGTGTTAAAAGTAGTTTTTGACAATGCGAATGGTAGGGAC ATCATTTTTTCGTCATTTCATCCGGATATTTGCATCATGCTTTCGTTAAAGCAACCGGTT ATTCCGATCTTGTTCCTGACGGAAGGTGGAAGTGAACAGATGGCCGATTTGAGAGCCTCA TCGTTACAAAACGGTATAAGATTTGCTAAGAAGTGGAATTTATTGGGGATTGTATCTGCT GCAGCACCCATTTTAAAGGCCCCACGGTTGGTGCAAGTGGTTAAGTCTAATGGGCTGGTA TGTGTCACTTACGGTGTGGACAACAATGATCCCGAAAATGCCAGTATACAAATTGAAGCA GGTGTAGATGCCGTGATCGTCGATAGTGTATTGGCAATTAGACGTGGGTTAACCAAAAAG AATGAAAAGTAA | |||||||||||||||||||||||||||
| Protein Properties | ||||||||||||||||||||||||||||
| Pfam Domain Function | ||||||||||||||||||||||||||||
| Protein Residues | 1223 | |||||||||||||||||||||||||||
| Protein Molecular Weight | 138013.0 | |||||||||||||||||||||||||||
| Protein Theoretical pI | 6.77 | |||||||||||||||||||||||||||
| Signalling Regions |
| |||||||||||||||||||||||||||
| Transmembrane Regions |
| |||||||||||||||||||||||||||
| Protein Sequence | >Glycerophosphodiester phosphodiesterase GDE1 MKFGKTFANHRIPEWSSQYVGYKSLKKMIKEITRLQEDIYRAHNKNSYDEGRPPTKMRDS SNSAQNYLDSPKIQKLLASFFFAVDRDIEKVDTFYNSQYAEYKKRFERLLSSNQFNEIKS TLVVDANKEDAVAQTLLTKDTREMNMLLKGTSQASRLSYHKDDLIEIQSILAELRKQFRN LKWYAELNKRAFGKILKKLDKKVGTNQQMSTMKTRILPLQFANDSLITKDLSLLKTIWEQ VTFRINSYERVMRSTSPNANANDNTEFFKIICVFIEEDDSKGLIRELTNLYSELSLIPTR IMISVLNKAALSKSLACIDAILKVIPSLNDSEDINRRNFFHHHIIAIGKLIRKQEILSRK KKSQPSKYTNSEGEIVTDLRTLHTTLSAPAESDSITEEEKSSACTLSYILEELPIHLRPC LFQHDNYKRTPLHYSCQYGLSEVTKLIIKLMKEWNIWNEIPIDDVSAFGDAESLTPLHLC VLGAHPKTTEVLLQSLDPNVKLKSSSLLHLATEWNNYPLLHVLLSSKRFDINYQDNELHE TPLYLACRLNFFEAAVCLLYNGADLEIREKLFGWTAIFVAAAEGFTDIVKLLIANNANFD IEDEGGWTPMEHAVLRGHLHIADMVQIRDELVTHPHSQLNSGSEEKEPLNEISAGELNER NENGNGGNKGSLGKLAGPIKSYGHRFLDNNESLILITLGSNDTRNKSPSISLSSEALAKV IGLETDCALSLVISCNDSIDKSSVILDLPLDDNVDAVDFKVPFKVDYSHTLYFDIVPTYG TRSLETHNRIDCQKNNNNYVMARGVSMLNKSYSSVGVNRSILNGSVTVPIIANHTLEILG TLKFEYIIITPFEHPQLPLERTETYWKSLVSTRVIGHRGLGKNNPNKSLQIGENTVESFI MAASLGASYVEFDVQLTKDNVPVVYHDFLVAETGVDIPMHELTLEQFLDLNNADKEHIQR GAGHSPHHVNGADTALQKYRGRSVDDSDVSTLRRAWDLHDNDPNGKSNNAHWSDNRMRLT KTFKKNNFKGNARGHSIASSFVTLKELFKKIPANVGFNIECKFPMLDEAEEEELGQIMME MNHWVDTVLKVVFDNANGRDIIFSSFHPDICIMLSLKQPVIPILFLTEGGSEQMADLRAS SLQNGIRFAKKWNLLGIVSAAAPILKAPRLVQVVKSNGLVCVTYGVDNNDPENASIQIEA GVDAVIVDSVLAIRRGLTKKNEK | |||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||
| External Links |
| |||||||||||||||||||||||||||
| General Reference |
| |||||||||||||||||||||||||||