| General Reference | - Churcher, C., Bowman, S., Badcock, K., Bankier, A., Brown, D., Chillingworth, T., Connor, R., Devlin, K., Gentles, S., Hamlin, N., Harris, D., Horsnell, T., Hunt, S., Jagels, K., Jones, M., Lye, G., Moule, S., Odell, C., Pearson, D., Rajandream, M., Rice, P., Rowley, N., Skelton, J., Smith, V., Barrell, B., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome IX." Nature 387:84-87.9169870
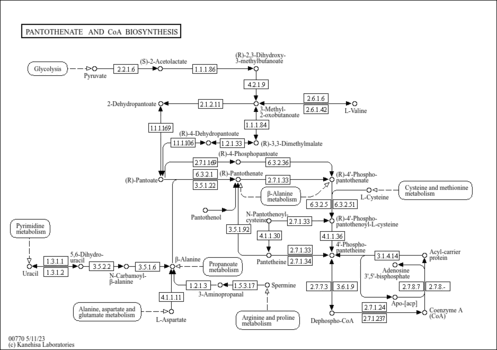
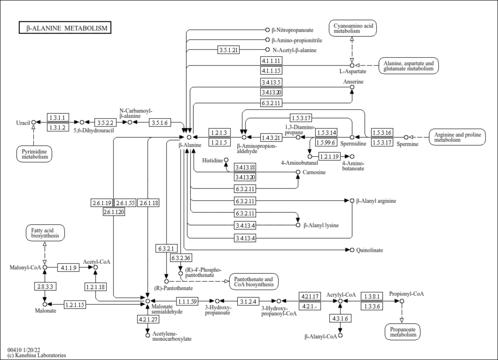
- White, W. H., Gunyuzlu, P. L., Toyn, J. H. (2001). "Saccharomyces cerevisiae is capable of de Novo pantothenic acid biosynthesis involving a novel pathway of beta-alanine production from spermine." J Biol Chem 276:10794-10800.11154694
- Kellis, M., Patterson, N., Endrizzi, M., Birren, B., Lander, E. S. (2003). "Sequencing and comparison of yeast species to identify genes and regulatory elements." Nature 423:241-254.12748633
- Huh, W. K., Falvo, J. V., Gerke, L. C., Carroll, A. S., Howson, R. W., Weissman, J. S., O'Shea, E. K. (2003). "Global analysis of protein localization in budding yeast." Nature 425:686-691.14562095
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
|
|---|