| Identification |
|---|
| Name | Putative pyridoxal kinase BUD16 |
|---|
| Synonyms | - Bud site selection protein 16
|
|---|
| Gene Name | BUD16 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in pyridoxal kinase activity |
|---|
| Specific Function | Required for synthesis of pyridoxal-5-phosphate from vitamin B6. Important for bud site selection |
|---|
| Cellular Location | Cytoplasm. Nucleus |
|---|
| SMPDB Pathways | Not Available |
|---|
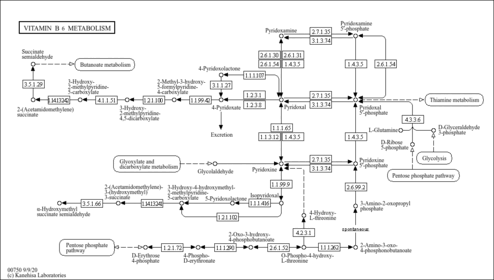
| KEGG Pathways | |
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | Not Available |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00109 | Adenosine triphosphate | Show | | YMDB00363 | Pyridoxal 5'-phosphate | Show | | YMDB00392 | Pyridoxal | Show | | YMDB00914 | ADP | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| transferase activity | | transferase activity, transferring phosphorus-containing groups | | kinase activity | | pyridoxal kinase activity | | catalytic activity | | Process |
|---|
| metabolic process | | small molecule metabolic process | | vitamin metabolic process | | water-soluble vitamin metabolic process | | vitamin B6 metabolic process | | pyridoxine metabolic process | | pyridoxine biosynthetic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 5 |
|---|
| Locus | YEL029C |
|---|
| Gene Sequence | >939 bp
ATGCCTCGTCTCTTGGCCACGCAGTCTCATGTTGTACATGGATATGTGGGAAATAAGGCT
GCAACGTTTCCCTTACAGTGTCTAGGCTGGGATGTGGATTGTTGTAACAGTGTTCAGTTT
TCCAACCATACCGGATATGGTTTAGATAAAGTGTTTGGGACTATAACGAGGGAAACAGAT
TTGAAGGAACTCTTATCAGGTCTTTTTGATAATTTTTCCCAAGACTACCAAGCCTTACTT
TCAGGCTACTTGCCCAATAAGAATTCCGTTCGATGTATGGGAACATACTACGCTAAATTT
AAAGAGGCGAATCCAGAAATGATATGGCTTATGGATCCTGTTATGGGTGACGAAGGACAA
TTATATGTTAGTGAGGATGTCATACCAGAATATAGAAAGCTGGCTTTGTCTCCAAAACAG
CTAGTCGATATAATAACTCCGAATCAATTTGAGCTGGAGATACTTTATGGGGGGGAAATC
AAGACTAAAGAACATTTGAAGAAGGCCTTAAAGAAATTACACCAAACTATCCCAGTGATC
ATTGTCACATCCTGTGACTGTAAGATGTTTGATGACAAAGATTTTATTTATTGCGTAGCT
TCAATGGAGGGTAAAACCCCCATTGTTTACAGAGTGCCGTTTATTGATTCGTATTTCACT
GGGGTTGGTGATTTGTTTTCTGCTTTGTTATTGGATCGAGTGTATAAGATCTTATCGAAC
CCTACAACAACATTAAAATTTGAAGACCAAGTCAATAATGTTCTTAATGTTATTCAGAAG
GTTTTGAAAATTACTAGGAGCTATGCTTCAGGAAAAATGAAAGCAAAAATGGGTTCCGCC
TTGGAAATGAAAGAAATGGAATTGCGTCTTATTGAATCAAGAGATATATATGAAACAATC
AACATTCATCAAACAGATTATATTTACGCAAGGTTGTGA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 312 |
|---|
| Protein Molecular Weight | 35558.89844 |
|---|
| Protein Theoretical pI | 6.51 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Putative pyridoxal kinase BUD16
MPRLLATQSHVVHGYVGNKAATFPLQCLGWDVDCCNSVQFSNHTGYGLDKVFGTITRETD
LKELLSGLFDNFSQDYQALLSGYLPNKNSVRCMGTYYAKFKEANPEMIWLMDPVMGDEGQ
LYVSEDVIPEYRKLALSPKQLVDIITPNQFELEILYGGEIKTKEHLKKALKKLHQTIPVI
IVTSCDCKMFDDKDFIYCVASMEGKTPIVYRVPFIDSYFTGVGDLFSALLLDRVYKILSN
PTTTLKFEDQVNNVLNVIQKVLKITRSYASGKMKAKMGSALEMKEMELRLIESRDIYETI
NIHQTDYIYARL |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Dietrich, F. S., Mulligan, J., Hennessy, K., Yelton, M. A., Allen, E., Araujo, R., Aviles, E., Berno, A., Brennan, T., Carpenter, J., Chen, E., Cherry, J. M., Chung, E., Duncan, M., Guzman, E., Hartzell, G., Hunicke-Smith, S., Hyman, R. W., Kayser, A., Komp, C., Lashkari, D., Lew, H., Lin, D., Mosedale, D., Davis, R. W., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome V." Nature 387:78-81.9169868
- Ni, L., Snyder, M. (2001). "A genomic study of the bipolar bud site selection pattern in Saccharomyces cerevisiae." Mol Biol Cell 12:2147-2170.11452010
- Huh, W. K., Falvo, J. V., Gerke, L. C., Carroll, A. S., Howson, R. W., Weissman, J. S., O'Shea, E. K. (2003). "Global analysis of protein localization in budding yeast." Nature 425:686-691.14562095
|
|---|