Long-chain-fatty-acid--CoA ligase 2 (P39518)
| Identification | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Long-chain-fatty-acid--CoA ligase 2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Name | FAA2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Enzyme Class | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General Function | Involved in catalytic activity | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Specific Function | Esterification, concomitant with transport, of endogenous long-chain fatty acids into metabolically active CoA thioesters for subsequent degradation or incorporation into phospholipids. Preferentially acts on C9:0-C13:0 fatty acids although C7:0-C17:0 fatty acids are tolerated | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Location | Cytoplasm. Mitochondrion. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
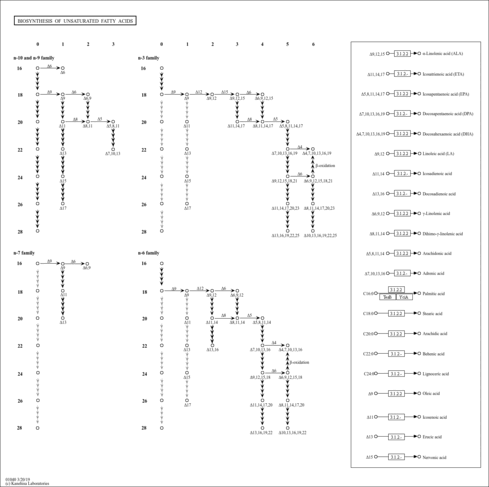
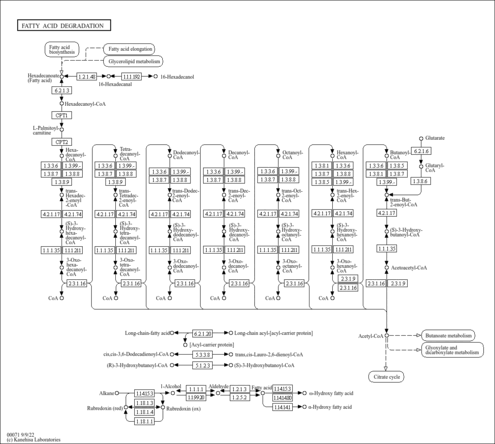
| KEGG Pathways |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Reactions |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Reactions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolites |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO Classification |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chromosome Location | chromosome 5 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Locus | YER015W | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Sequence | >2235 bp ATGGCCGCTCCAGATTATGCACTTACCGATTTAATTGAATCGGATCCTCGTTTCGAAAGT TTGAAGACAAGATTAGCCGGTTACACCAAAGGCTCTGATGAATATATTGAAGAGCTATAC TCTCAATTACCACTGACCAGCTATCCCAGGTACAAAACATTTTTAAAGAAACAGGCGGTT GCCATTTCGAATCCGGATAATGAAGCTGGTTTTAGCTCGATTTATAGGAGTTCTCTTTCT TCTGAAAATCTAGTGAGCTGTGTGGATAAAAACTTAAGAACTGCATACGATCACTTCATG TTTTCTGCAAGGAGATGGCCTCAACGTGACTGTTTAGGTTCAAGGCCAATTGATAAAGCC ACAGGCACCTGGGAGGAAACATTCCGTTTCGAGTCGTACTCCACGGTATCTAAAAGATGT CATAATATCGGAAGTGGTATATTGTCTTTGGTAAACACGAAAAGGAAACGTCCTTTGGAA GCCAATGATTTTGTTGTTGCTATCTTATCACACAACAACCCTGAATGGATCCTAACAGAT TTGGCCTGTCAGGCCTATTCTCTAACTAACACGGCTTTGTACGAAACATTAGGTCCAAAC ACCTCCGAGTACATATTGAATTTAACCGAGGCCCCCATTCTGATTTTTGCAAAATCAAAT ATGTATCATGTATTGAAGATGGTGCCTGATATGAAATTTGTTAATACTTTGGTTTGTATG GATGAATTAACTCATGACGAGCTCCGTATGCTAAATGAATCGTTGCTACCCGTTAAGTGC AACTCTCTCAATGAAAAAATCACATTTTTTTCATTGGAGCAGGTAGAACAAGTTGGTTGC TTTAACAAAATTCCTGCAATTCCACCTACCCCAGATTCCTTGTATACTATTTCGTTTACT TCTGGTACTACAGGTTTACCTAAAGGTGTGGAAATGTCTCACAGAAACATTGCGTCTGGG ATAGCATTTGCTTTTTCTACCTTCAGAATACCGCCAGATAAAAGAAACCAACAGTTATAT GATATGTGTTTTTTGCCATTGGCTCATATTTTTGAAAGAATGGTTATTGCGTATGATCTA GCCATCGGGTTTGGAATAGGCTTCTTACATAAACCAGACCCAACTGTATTGGTAGAGGAT TTGAAGATTTTGAAACCTTACGCGGTTGCCCTGGTTCCTAGAATATTAACACGGTTTGAA GCCGGTATAAAAAACGCTTTGGATAAATCGACTGTCCAGAGGAACGTAGCAAATACTATA TTGGATTCTAAATCGGCCAGATTTACCGCAAGAGGTGGTCCAGATAAATCGATTATGAAT TTTCTAGTTTATCATCGCGTATTGATTGATAAAATCAGAGACTCTTTAGGTTTGTCCAAT AACTCGTTTATAATTACCGGATCAGCTCCCATATCTAAAGATACCTTACTATTTTTAAGA AGTGCCTTGGATATTGGTATAAGACAGGGCTACGGCTTAACTGAAACTTTTGCTGGTGTC TGTTTAAGCGAACCGTTTGAAAAAGATGTCGGATCTTGTGGTGCCATAGGTATTTCTGCA GAATGTAGATTGAAGTCTGTTCCAGAAATGGGTTACCATGCCGACAAGGATTTAAAAGGT GAACTGCAAATTCGTGGCCCACAGGTTTTTGAAAGATATTTTAAAAATCCGAATGAAACT TCAAAAGCCGTTGACCAAGATGGTTGGTTTTCCACGGGAGATGTTGCATTTATCGATGGA AAAGGTCGCATCAGCGTCATTGATCGAGTCAAGAACTTTTTCAAGCTAGCACATGGTGAA TATATTGCTCCAGAGAAAATCGAAAATATTTATTTATCATCATGCCCCTATATCACGCAA ATATTTGTCTTTGGAGATCCTTTAAAGACATTTTTAGTTGGCATCGTTGGTGTTGATGTT GATGCAGCGCAACCGATTTTAGCTGCAAAGCACCCAGAGGTGAAAACGTGGACTAAGGAA GTGCTAGTAGAAAACTTAAATCGTAATAAAAAGCTAAGGAAGGAATTTTTAAACAAAATT AATAAATGCACCGATGGGCTACAAGGATTCGAAAAATTGCATAACATCAAAGTCGGACTT GAGCCTTTAACTCTCGAGGATGATGTTGTGACGCCAACTTTTAAAATAAAGCGTGCCAAA GCATCAAAATTCTTCAAAGATACATTAGACCAACTATACGCCGAAGGTTCACTAGTCAAG ACAGAAAAGCTTTAG | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pfam Domain Function |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Residues | 744 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Molecular Weight | 83437.10156 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Theoretical pI | 7.82 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Signalling Regions |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Transmembrane Regions |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence | >Long-chain-fatty-acid--CoA ligase 2 MAAPDYALTDLIESDPRFESLKTRLAGYTKGSDEYIEELYSQLPLTSYPRYKTFLKKQAV AISNPDNEAGFSSIYRSSLSSENLVSCVDKNLRTAYDHFMFSARRWPQRDCLGSRPIDKA TGTWEETFRFESYSTVSKRCHNIGSGILSLVNTKRKRPLEANDFVVAILSHNNPEWILTD LACQAYSLTNTALYETLGPNTSEYILNLTEAPILIFAKSNMYHVLKMVPDMKFVNTLVCM DELTHDELRMLNESLLPVKCNSLNEKITFFSLEQVEQVGCFNKIPAIPPTPDSLYTISFT SGTTGLPKGVEMSHRNIASGIAFAFSTFRIPPDKRNQQLYDMCFLPLAHIFERMVIAYDL AIGFGIGFLHKPDPTVLVEDLKILKPYAVALVPRILTRFEAGIKNALDKSTVQRNVANTI LDSKSARFTARGGPDKSIMNFLVYHRVLIDKIRDSLGLSNNSFIITGSAPISKDTLLFLR SALDIGIRQGYGLTETFAGVCLSEPFEKDVGSCGAIGISAECRLKSVPEMGYHADKDLKG ELQIRGPQVFERYFKNPNETSKAVDQDGWFSTGDVAFIDGKGRISVIDRVKNFFKLAHGE YIAPEKIENIYLSSCPYITQIFVFGDPLKTFLVGIVGVDVDAAQPILAAKHPEVKTWTKE VLVENLNRNKKLRKEFLNKINKCTDGLQGFEKLHNIKVGLEPLTLEDDVVTPTFKIKRAK ASKFFKDTLDQLYAEGSLVKTEKL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General Reference |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||