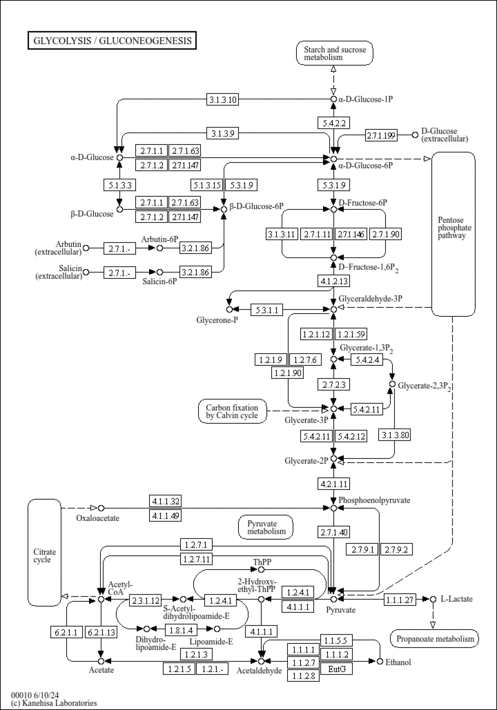
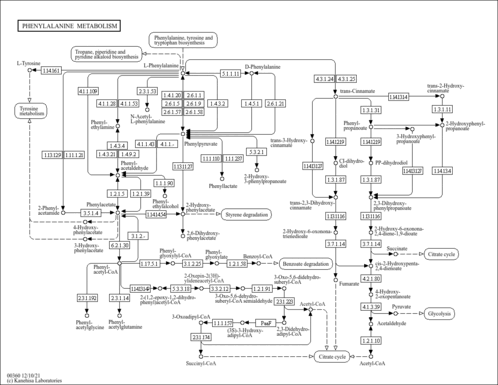
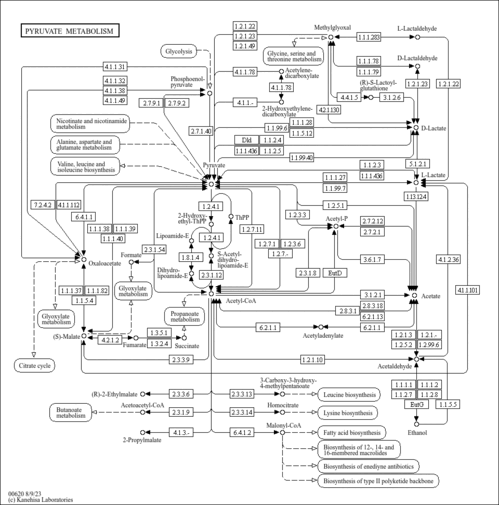
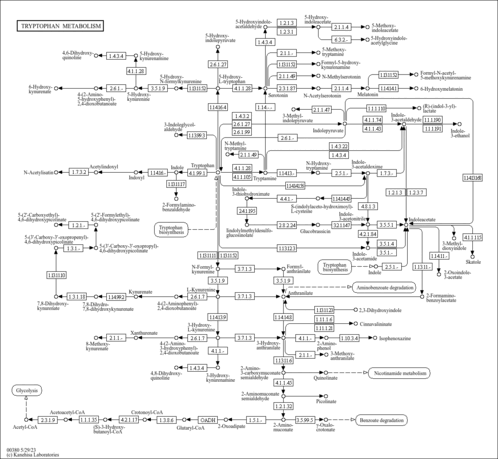
| General Reference | - Hohmann, S. (1991). "Characterization of PDC6, a third structural gene for pyruvate decarboxylase in Saccharomyces cerevisiae." J Bacteriol 173:7963-7969.1744053
- Tettelin, H., Agostoni Carbone, M. L., Albermann, K., Albers, M., Arroyo, J., Backes, U., Barreiros, T., Bertani, I., Bjourson, A. J., Bruckner, M., Bruschi, C. V., Carignani, G., Castagnoli, L., Cerdan, E., Clemente, M. L., Coblenz, A., Coglievina, M., Coissac, E., Defoor, E., Del Bino, S., Delius, H., Delneri, D., de Wergifosse, P., Dujon, B., Kleine, K., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome VII." Nature 387:81-84.9169869
- Dickinson, J. R., Lanterman, M. M., Danner, D. J., Pearson, B. M., Sanz, P., Harrison, S. J., Hewlins, M. J. (1997). "A 13C nuclear magnetic resonance investigation of the metabolism of leucine to isoamyl alcohol in Saccharomyces cerevisiae." J Biol Chem 272:26871-26878.9341119
- Dickinson, J. R., Harrison, S. J., Hewlins, M. J. (1998). "An investigation of the metabolism of valine to isobutyl alcohol in Saccharomyces cerevisiae." J Biol Chem 273:25751-25756.9748245
- Flikweert, M. T., de Swaaf, M., van Dijken, J. P., Pronk, J. T. (1999). "Growth requirements of pyruvate-decarboxylase-negative Saccharomyces cerevisiae." FEMS Microbiol Lett 174:73-79.10234824
- Dickinson, J. R., Harrison, S. J., Dickinson, J. A., Hewlins, M. J. (2000). "An investigation of the metabolism of isoleucine to active Amyl alcohol in Saccharomyces cerevisiae." J Biol Chem 275:10937-10942.10753893
- Neuser, F., Zorn, H., Berger, R. G. (2000). "Generation of odorous acyloins by yeast pyruvate decarboxylases and their occurrence in sherry and soy sauce." J Agric Food Chem 48:6191-6195.11141278
- Dickinson, J. R., Salgado, L. E., Hewlins, M. J. (2003). "The catabolism of amino acids to long chain and complex alcohols in Saccharomyces cerevisiae." J Biol Chem 278:8028-8034.12499363
- Vuralhan, Z., Morais, M. A., Tai, S. L., Piper, M. D., Pronk, J. T. (2003). "Identification and characterization of phenylpyruvate decarboxylase genes in Saccharomyces cerevisiae." Appl Environ Microbiol 69:4534-4541.12902239
- Iding, H., Siegert, P., Mesch, K., Pohl, M. (1998). "Application of alpha-keto acid decarboxylases in biotransformations." Biochim Biophys Acta 1385:307-322.9655924
- Hohmann, S., Meacock, P. A. (1998). "Thiamin metabolism and thiamin diphosphate-dependent enzymes in the yeast Saccharomyces cerevisiae: genetic regulation." Biochim Biophys Acta 1385:201-219.9655908
- Huh, W. K., Falvo, J. V., Gerke, L. C., Carroll, A. S., Howson, R. W., Weissman, J. S., O'Shea, E. K. (2003). "Global analysis of protein localization in budding yeast." Nature 425:686-691.14562095
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
- Albuquerque, C. P., Smolka, M. B., Payne, S. H., Bafna, V., Eng, J., Zhou, H. (2008). "A multidimensional chromatography technology for in-depth phosphoproteome analysis." Mol Cell Proteomics 7:1389-1396.18407956
|
|---|