| Identification |
|---|
| Name | Ectonucleotide pyrophosphatase/phosphodiesterase 1 |
|---|
| Synonyms | - E-NPP 1
- Alkaline phosphodiesterase 1
- Nucleotide pyrophosphatase
- NPPase
|
|---|
| Gene Name | NPP1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in catalytic activity |
|---|
| Specific Function | Mediates extracellular nucleotide derived phosphate hydrolysis along with NPP2 and PHO5 |
|---|
| Cellular Location | Membrane; Single-pass type II membrane protein (Potential) |
|---|
| SMPDB Pathways | |
|---|
| KEGG Pathways | | Nicotinate and nicotinamide metabolism | ec00760 | 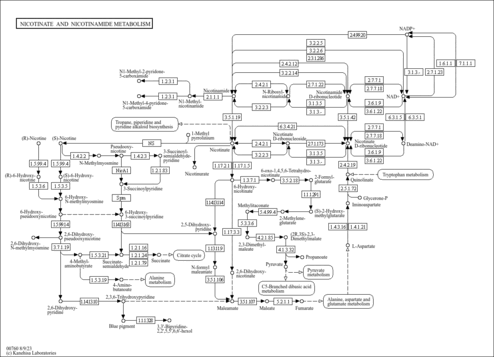 | | Pantothenate and CoA biosynthesis | ec00770 | 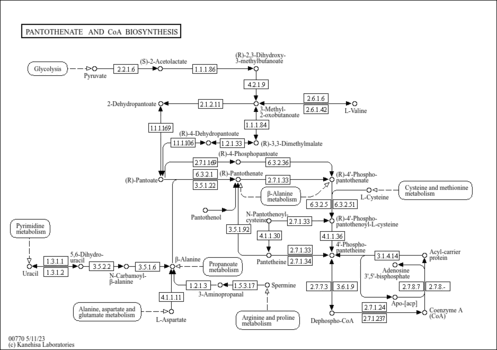 | | Purine metabolism | ec00230 |  | | Riboflavin metabolism | ec00740 | 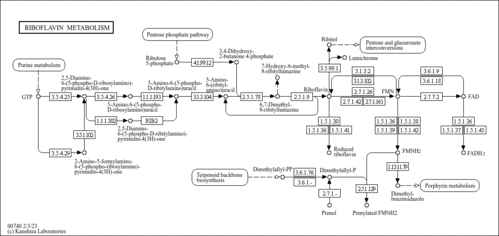 | | Starch and sucrose metabolism | ec00500 | 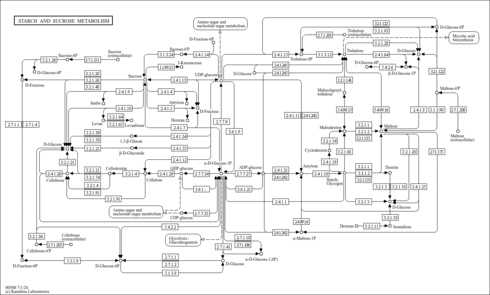 |
|
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | Not Available |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00049 | Uridine 5'-monophosphate | Show | | YMDB00261 | Guanosine monophosphate | Show | | YMDB00288 | Glucose 1-phosphate | Show | | YMDB00386 | D-Mannose 1-phosphate | Show | | YMDB00415 | UDP-D-glucose | Show | | YMDB00890 | water | Show |
|
|---|
| GO Classification | | Component |
|---|
| Not Available | | Function |
|---|
| catalytic activity | | Process |
|---|
| metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 3 |
|---|
| Locus | YCR026C |
|---|
| Gene Sequence | >2229 bp
ATGGAACTTCAGAATGATTTAGAGTCGCTCGATAACGAGCTGAATGATTTTAGTGAAGAT
CCATTTCGTGATGATTTCATAACGGATGAAGATGCTGTAAGATCGGGGTGGCGATCTGCG
TGGACCAGGATGAAATATTGGTTTTATAAGAATAGACTGAAGTGGACAAACAATCCCATA
GTGATTGGCGACGCGAAAGATAGTAGGGATGGTTCTAACTTTAGAAGGGGTATACCGCTA
TATGAATTAGACGCGAATGGTCAACCCATTGATACTGAACTTGTTGATGAGAATGAACTT
TCTTTTGGAACGGGATTTCATTCCAAAGTGCCTTTTAAAATAATATTTCGCACATTGTTT
GGCTCGCTGGTGTTTGCCATTTTTTTAATTCTGATGATTAACATAGCAAAACCCCATCAC
TCCACGAGAGTGCTATCGCACTTTGGCAGTCCTGAATTTGACCCTTACGTGAAGTATTTT
AACGGTACGCATGAATTTTTCCCCTTAACGATAGTAATTTCACTAGACGGTTTCCATCCT
TCACTCATATCTAAGAGGAACACACCGTTTTTACATGACTTATATGAATTGAAATATGAT
GGAGGTATGAATATCACGTCCACACCTTTTATGGTACCCAGCTTCCCTACGGAGACCTTT
CCCAACCATTGGACGTTGGTTACTGGACAATACCCAATACACCACGGTATAGTCTCTAAC
GTATTTTGGGATCCTGATCTTAATGAGGAATTCCATCCAGGTGTATTGGACCCTCGAATA
TGGAACAATAATGATACAGAACCAATATGGCAAACTGTTCAGTCTGCATTTGACGGTGAT
ATACCATTCAAAGCTGCTACCCATATGTGGCCAGGTAGCGATGTGAATTATACCAAGTAT
AATGAAGAGAAACTACAACCTGAACATAAAAATCCTATTGCTAGAGAGAGAACTCCATTT
TACTTCGACGAATTCAATGCTAAAGAACCACTTTCGCAAAAATTATCCAAGATTATTGAA
TATGTGGATATGAGTACACTGAACGAAAGACCACAGCTAATTCTCGGTTATGTACCGAAC
GTAGATGCCTTTGGACATAAGCATGGATATCCGTCAGAGTCGGAATACTATTATGAAGAC
TTCACTGAAACACTGGGGGAAGTAGATACATTTCTGAAGCAACTAGTGGAATCGCTGCAA
GAAAGAAATTTAACCAGCTTTACTAATTTGGTCATTGTTAGCGATCATGGTATGAGCGAT
ATCGTAGTTCCCTCAAATGTTATTATATGGGAAGACTTACTGGACGAAAAATTGAGGAAG
GATTATGTATCGCACGCATATCTAGAGGGTCCGATGATGGCTATATCGTTGAAAGATTCC
GGAAACATCAATGAGGTTTACCACAATTTAAAGACTTCTATAGATGAAGACAAGTATACG
GTTTACGTTAATGGAAATTTCCCCAAAGAATGGAACTTTAATGATGGAAAAAATCATCAC
ATGGCGTCAATCTGGATTGTGCCCGAGCCTGGGTATGCAGTGATGAAGAAAGAACAATTG
AAGAAGGTGGCAAAAGGTGATCATAAGGACAAAAACGAAGACAATGTGTTCACGATTGGA
TCACATGGATACGACAATAACGCGATCGATATGAGATCTGTATTTATTGGTATGGGGCCA
TATTTTCCACAGGGATACATTGAGCCGTTCCAAAATACCGAAATTTACAACCTTTTGTGT
GATATTTGCGGTGTGGCAGAAAAGGACAGAAATTCCAATGATGGGACTGGGATGCTTATG
AACCAACTCCGCGAACCCCAGAGCAGCGAAGAAGTAGAGATTGAAGATGACTTTGATTAT
TTGGTCAGTAAGTTTGGTGAATTCAGCACTTATAATATAATTTGGGGCGGGTACCCCGAA
GAGACAGAACAAGACAATGTTGACAATGATAATGATGACAACGACGATGGAAACACTGAT
GAAATAGCCGCTATGCCATCTTCGTCATTAACGATAAAACTAGAAATGACAACTTCAATA
CCATCAGCAACTGAGACTCTACTGGGCGAAACATCACCATCATCAAGAAGCAGCAGCAGC
AGCAGCATACAAGCTAGCGCTACTGCTAGCACAGTGGGGGATTGGCTTCAAGACATAATC
AACGACGCAAAAGATCTCATTGACGACATAATTGACAGCATCGACGATTTAGTCGATTCT
GATACCTAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 742 |
|---|
| Protein Molecular Weight | 84733.20313 |
|---|
| Protein Theoretical pI | 4.24 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Ectonucleotide pyrophosphatase/phosphodiesterase 1
MELQNDLESLDNELNDFSEDPFRDDFITDEDAVRSGWRSAWTRMKYWFYKNRLKWTNNPI
VIGDAKDSRDGSNFRRGIPLYELDANGQPIDTELVDENELSFGTGFHSKVPFKIIFRTLF
GSLVFAIFLILMINIAKPHHSTRVLSHFGSPEFDPYVKYFNGTHEFFPLTIVISLDGFHP
SLISKRNTPFLHDLYELKYDGGMNITSTPFMVPSFPTETFPNHWTLVTGQYPIHHGIVSN
VFWDPDLNEEFHPGVLDPRIWNNNDTEPIWQTVQSAFDGDIPFKAATHMWPGSDVNYTKY
NEEKLQPEHKNPIARERTPFYFDEFNAKEPLSQKLSKIIEYVDMSTLNERPQLILGYVPN
VDAFGHKHGYPSESEYYYEDFTETLGEVDTFLKQLVESLQERNLTSFTNLVIVSDHGMSD
IVVPSNVIIWEDLLDEKLRKDYVSHAYLEGPMMAISLKDSGNINEVYHNLKTSIDEDKYT
VYVNGNFPKEWNFNDGKNHHMASIWIVPEPGYAVMKKEQLKKVAKGDHKDKNEDNVFTIG
SHGYDNNAIDMRSVFIGMGPYFPQGYIEPFQNTEIYNLLCDICGVAEKDRNSNDGTGMLM
NQLREPQSSEEVEIEDDFDYLVSKFGEFSTYNIIWGGYPEETEQDNVDNDNDDNDDGNTD
EIAAMPSSSLTIKLEMTTSIPSATETLLGETSPSSRSSSSSSIQASATASTVGDWLQDII
NDAKDLIDDIIDSIDDLVDSDT |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Oliver, S. G., van der Aart, Q. J., Agostoni-Carbone, M. L., Aigle, M., Alberghina, L., Alexandraki, D., Antoine, G., Anwar, R., Ballesta, J. P., Benit, P., et, a. l. .. (1992). "The complete DNA sequence of yeast chromosome III." Nature 357:38-46.1574125
- Bolle, P. A., Gilliquet, V., Berben, G., Dumont, J., Hilger, F. (1992). "The complete sequence of K3B, a 7.9 kb fragment between PGK1 and CRY1 on chromosome III, reveals the presence of seven open reading frames." Yeast 8:205-213.1574926
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
- Kennedy, E. J., Pillus, L., Ghosh, G. (2005). "Pho5p and newly identified nucleotide pyrophosphatases/ phosphodiesterases regulate extracellular nucleotide phosphate metabolism in Saccharomyces cerevisiae." Eukaryot Cell 4:1892-1901.16278456
- Albuquerque, C. P., Smolka, M. B., Payne, S. H., Bafna, V., Eng, J., Zhou, H. (2008). "A multidimensional chromatography technology for in-depth phosphoproteome analysis." Mol Cell Proteomics 7:1389-1396.18407956
- Kung, L. A., Tao, S. C., Qian, J., Smith, M. G., Snyder, M., Zhu, H. (2009). "Global analysis of the glycoproteome in Saccharomyces cerevisiae reveals new roles for protein glycosylation in eukaryotes." Mol Syst Biol 5:308.19756047
|
|---|




