Ribonucleoside-diphosphate reductase large chain 2 (P21672)
| Identification | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Ribonucleoside-diphosphate reductase large chain 2 | ||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||
| Gene Name | RNR3 | ||||||||||||||||||||||||
| Enzyme Class | |||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||
| General Function | Involved in oxidation-reduction process | ||||||||||||||||||||||||
| Specific Function | Provides the precursors necessary for DNA synthesis. Catalyzes the biosynthesis of deoxyribonucleotides from the corresponding ribonucleotides | ||||||||||||||||||||||||
| Cellular Location | Cytoplasm | ||||||||||||||||||||||||
| SMPDB Pathways |
| ||||||||||||||||||||||||
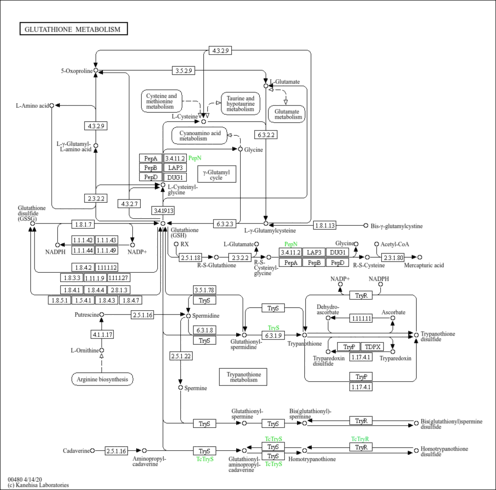
| KEGG Pathways |
| ||||||||||||||||||||||||
| SMPDB Reactions | Not Available | ||||||||||||||||||||||||
| KEGG Reactions | Not Available | ||||||||||||||||||||||||
| Metabolites |
| ||||||||||||||||||||||||
| GO Classification |
| ||||||||||||||||||||||||
| Gene Properties | |||||||||||||||||||||||||
| Chromosome Location | Not Available | ||||||||||||||||||||||||
| Locus | |||||||||||||||||||||||||
| Gene Sequence | >YIL066C RNR3 SGDID:S000001328, Chr IX from 240708-238099, reverse complement, Verified ORF, "Minor isoform of the large subunit of ribonucleotide-diphosphate reductase; the RNR complex catalyzes rate-limiting step in dNTP synthesis, regulated by DNA repli ATGTACGTTATTAAAAGAGACGGCCGCAAAGAGCCCGTTCAATTCGATAAAATTACCTCC CGTATCACCCGTTTGTCATACGGTTTAGACCCAAACCGTATTGATGCTGTTAAGGTAACC CAACGTATTATTTCTGGTGTGTACTCCGGTGTTACTACCGTTGAGCTGGACAATCTTGCA GCTGAAACATGTGCATACATGACCACTGTGCACCCTGATTATGCCACTCTAGCCGCTAGA ATCGCCATCTCTAACTTACATAAGCAAACCACAAAGCAATTCTCCAAAGTTATTGAGGAT TTACACGACTGGATTAACCCAGCTACTGGAAAGCATGCTCCTATGATTTCGGACGAAATT TACAACATTGTCATGGAAAACAAAGATACTTTGAACTCGGCCATCGTGTACGATAGGGAT TTCCAGTATACGTATTTCGGATTCAAGACACTGGAGCGTTCGTACTTGCTAAGACTGAAC GGTGAAGTGGCAGAACGTCCTCAGCATTTGGTAATGCGTGTGGCGCTAGGTATCCATGGT AGCGATATCGAATCTGTGCTGAAGACTTATAATTTGATGTCGTTAAGATACTTCACTCAC GCTTCCCCAACTTTATTCAACGCTGGTACGCCACATCCTCAAATGTCTTCATGTTTCTTA ATTGCCATGAAGGATGACTCTATCGAAGGTATTTATGATACTTTGAAAGAATGTGCTATG ATTTCCAAAACTGCAGGTGGTGTTGGTCTTCATATCAACAACATCCGTTCCACAGGTTCT TATATCGCTGGTACCAACGGTACTTCAAACGGGTTGATTCCTATGATTCGTGTTTTCAAT AATACTGCCCGTTATGTGGACCAGGGTGGTAACAAGAGACCTGGTGCTTTCGCCCTTTTC TTGGAGCCATGGCATGCAGATATCTTCGACTTTGTCGATATCAGAAAAACACATGGTAAG GAAGAAATTCGTGCAAGAGATTTGTTCCCTGCTCTATGGATCCCTGATCTTTTCATGAAA CGTGTTCAAGAGGATGGGCCTTGGACTTTGTTTTCGCCCAGTGCTGCCCCAGGTTTAGAT GATGTGTGGGGTGATGAATTTGAAGAACTATATACTCGTTACGAAAGAGAAGGTCGTGGT AAAACAATTAAAGCCCAAAAGTTGTGGTATGCCATTTTGCAAGCACAGACAGAAACAGGT ACACCTTTCATGGTTTATAAGGACGCATGTAACAGGAAGACAAACCAACAGAACTTAGGT ACTATCAAATCATCTAATTTATGTTGTGAAATCGTCGAATATTCCTCCCCGGATGAAACT GCAGTTTGTAATTTAGCTTCTATTGCCCTACCAGCATTCGTTGAGGTTTCAGAAGATGGT AAAACTGCAAGCTATAATTTCGAGAGATTACACGAGATTGCTAAAGTCATTACTCACAAC TTGAACAGAGTTATCGACCGTAATTACTATCCAGTTCCCGAGGCTAGAAATTCAAATATG AAGCATAGACCTATTGCTCTTGGTGTCCAGGGTTTGGCCGATACTTATATGATGTTGCGT CTACCCTTTGAATCGGAAGAAGCTCAAACTCTAAACAAACAAATCTTCGAAACTATTTAC CATGCTACTCTTGAAGCCTCCTGTGAATTGGCCCAAAAAGAAGGTAAATATTCTACTTTT GAAGGTTCTCCAGCTTCTAAGGGTATTTTACAATTCGATATGTGGAACGCTAAACCATTT GGCATGTGGGATTGGGAAACCTTAAGAAAGGACATTGTTAAACATGGGTTAAGAAACTCT TTGACTATGGCACCAATGCCAACCGCCTCAACTTCCCAAATTCTTGGTTATAATGAATGC TTCGAACCAGTGACCTCAAACATGTACTCTCGTCGTGTCCTGTCTGGTGAATTCCAAGTT GTTAATCCATATTTACTACGTGATTTAGTCGACCTGGGTATTTGGGATGATAGTATGAAA CAATATCTAATTACACAAAATGGTTCTATTCAAGGCTTACCAAATGTGCCACAAGAATTG AAGGAATTATACAAAACCGTCTGGGAAATCTCTCAAAAGACCATTATCAATATGGCTGCT GATCGTGCCATCTACATCGATCAGTCTCATTCCTTGAATCTTTTCTTGCAAGCACCATCA ATGGGTAAGATTACTAGTATGCATTTCTACGGTTGGAAGAAGGGTTTAAAAACTGGTATG TACTACTTAAGAACGCAAGCCGCCTCCGCTGCTATTCAATTTACCATTGATCAAGAGGTT GCCGATCAAGCCGCTACACATATTGCTTCCGTCTCAGAATTGGATCGTCCAGTTTATGTT CCAAAGGGTACAAAATTCTCTGAACAAAAGGCGGCATCTGCGCTTACCGAAAGCTCAGAT AATGAGAAGGATGCATCTCCAGTTCCATCCGAACAATCATCGGTGTCGAGTGCCATGTCA AATGTGAAATTGGAAGATAGTGTTGCCCCAGCAGTTCCAACGGAAACAATAAAAGAAGAT TCCGACGAGAAGAAATGTGACATTTACAATGAAAAGGTGATTGCTTGTACTGCTCCTACT CCAGAAGCTTGTGAGTCATGTTCCGGTTGA | ||||||||||||||||||||||||
| Protein Properties | |||||||||||||||||||||||||
| Pfam Domain Function | |||||||||||||||||||||||||
| Protein Residues | 869 | ||||||||||||||||||||||||
| Protein Molecular Weight | 97513.79688 | ||||||||||||||||||||||||
| Protein Theoretical pI | 5.86 | ||||||||||||||||||||||||
| Signalling Regions |
| ||||||||||||||||||||||||
| Transmembrane Regions |
| ||||||||||||||||||||||||
| Protein Sequence | >Ribonucleoside-diphosphate reductase large chain 2 MYVIKRDGRKEPVQFDKITSRITRLSYGLDPNRIDAVKVTQRIISGVYSGVTTVELDNLA AETCAYMTTVHPDYATLAARIAISNLHKQTTKQFSKVIEDLHDWINPATGKHAPMISDEI YNIVMENKDTLNSAIVYDRDFQYTYFGFKTLERSYLLRLNGEVAERPQHLVMRVALGIHG SDIESVLKTYNLMSLRYFTHASPTLFNAGTPHPQMSSCFLIAMKDDSIEGIYDTLKECAM ISKTAGGVGLHINNIRSTGSYIAGTNGTSNGLIPMIRVFNNTARYVDQGGNKRPGAFALF LEPWHADIFDFVDIRKTHGKEEIRARDLFPALWIPDLFMKRVQEDGPWTLFSPSAAPGLD DVWGDEFEELYTRYEREGRGKTIKAQKLWYAILQAQTETGTPFMVYKDACNRKTNQQNLG TIKSSNLCCEIVEYSSPDETAVCNLASIALPAFVEVSEDGKTASYNFERLHEIAKVITHN LNRVIDRNYYPVPEARNSNMKHRPIALGVQGLADTYMMLRLPFESEEAQTLNKQIFETIY HATLEASCELAQKEGKYSTFEGSPASKGILQFDMWNAKPFGMWDWETLRKDIVKHGLRNS LTMAPMPTASTSQILGYNECFEPVTSNMYSRRVLSGEFQVVNPYLLRDLVDLGIWDDSMK QYLITQNGSIQGLPNVPQELKELYKTVWEISQKTIINMAADRAIYIDQSHSLNLFLQAPS MGKITSMHFYGWKKGLKTGMYYLRTQAASAAIQFTIDQEVADQAATHIASVSELDRPVYV PKGTKFSEQKAASALTESSDNEKDASPVPSEQSSVSSAMSNVKLEDSVAPAVPTETIKED SDEKKCDIYNEKVIACTAPTPEACESCSG | ||||||||||||||||||||||||
| References | |||||||||||||||||||||||||
| External Links |
| ||||||||||||||||||||||||
| General Reference |
| ||||||||||||||||||||||||