| Identification |
|---|
| Name | Pyruvate carboxylase 1 |
|---|
| Synonyms | - Pyruvic carboxylase 1
- PCB 1
|
|---|
| Gene Name | PYC1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in catalytic activity |
|---|
| Specific Function | Pyruvate carboxylase catalyzes a 2-step reaction, involving the ATP-dependent carboxylation of the covalently attached biotin in the first step and the transfer of the carboxyl group to pyruvate in the second |
|---|
| Cellular Location | Cytoplasm |
|---|
| SMPDB Pathways | |
|---|
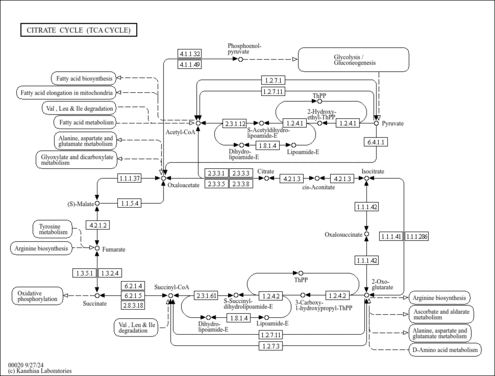
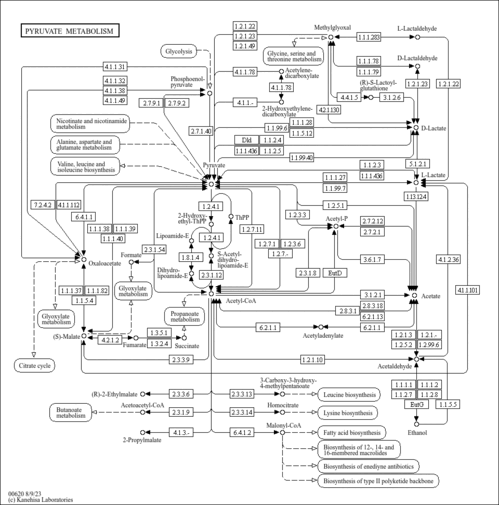
| KEGG Pathways | |
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00109 | Adenosine triphosphate | Show | | YMDB00175 | Pyruvic acid | Show | | YMDB00235 | Oxalacetic acid | Show | | YMDB00382 | Carbonic acid | Show | | YMDB00862 | hydron | Show | | YMDB00907 | phosphate | Show | | YMDB00914 | ADP | Show | | YMDB16118 | Hydrogen carbonate | Show |
|
|---|
| GO Classification | | Component |
|---|
| cell part | | intracellular part | | cytoplasm | | Function |
|---|
| catalytic activity | | ligase activity | | binding | | ligase activity, forming carbon-carbon bonds | | pyruvate carboxylase activity | | nucleoside binding | | purine nucleoside binding | | adenyl nucleotide binding | | adenyl ribonucleotide binding | | ATP binding | | vitamin binding | | biotin binding | | Process |
|---|
| monosaccharide metabolic process | | metabolic process | | hexose metabolic process | | glucose metabolic process | | gluconeogenesis | | small molecule metabolic process | | alcohol metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 7 |
|---|
| Locus | YGL062W |
|---|
| Gene Sequence | >3537 bp
ATGTCGCAAAGAAAATTCGCCGGCTTGAGAGATAACTTCAATCTCTTGGGTGAAAAGAAC
AAAATATTGGTGGCTAATAGAGGAGAAATTCCAATCAGAATTTTTCGTACCGCTCATGAA
CTGTCTATGCAGACGGTAGCTATATATTCTCATGAAGATCGTCTTTCAACGCACAAACAA
AAGGCTGACGAAGCATACGTCATAGGTGAAGTAGGCCAATATACCCCCGTCGGCGCTTAT
TTGGCCATTGACGAAATCATTTCCATTGCCCAAAAACACCAGGTAGATTTCATCCATCCA
GGTTATGGGTTCTTGTCTGAAAATTCGGAATTTGCCGACAAAGTAGTGAAGGCCGGTATC
ACTTGGATTGGCCCTCCAGCTGAAGTTATTGACTCCGTGGGTGATAAGGTCTCAGCTAGA
AACCTGGCAGCAAAAGCTAATGTGCCCACCGTTCCTGGTACACCAGGTCCTATAGAAACT
GTAGAGGAAGCACTTGACTTCGTCAATGAATACGGCTACCCGGTGATCATTAAGGCCGCC
TTTGGTGGTGGTGGTAGAGGTATGAGAGTCGTTAGAGAAGGTGACGACGTGGCAGATGCC
TTTCAACGTGCTACCTCCGAAGCCCGTACTGCCTTCGGTAATGGTACCTGCTTTGTGGAA
AGATTCTTGGACAAGCCAAAGCATATTGAAGTTCAATTGTTGGCCGATAACCACGGAAAC
GTGGTTCATCTTTTCGAAAGAGACTGTTCCGTGCAGAGAAGACACCAAAAGGTTGTCGAA
GTGGCCCCAGCAAAGACTTTACCCCGTGAAGTCCGTGACGCCATTTTGACAGATGCAGTT
AAATTGGCCAAAGAGTGTGGCTACAGAAATGCGGGTACTGCTGAATTCTTGGTTGATAAC
CAAAATAGACACTATTTCATTGAAATTAATCCAAGAATCCAAGTGGAACATACCATCACA
GAAGAAATTACCGGTATAGATATTGTGGCGGCTCAGATCCAAATTGCGGCAGGTGCCTCT
CTACCCCAGCTGGGCCTATTCCAGGACAAAATTACGACTCGTGGCTTTGCCATTCAGTGC
CGTATTACCACGGAAGACCCTGCTAAGAACTTCCAACCAGATACCGGTAGAATAGAAGTG
TACCGTTCTGCAGGTGGTAATGGTGTTAGACTGGATGGTGGTAACGCCTATGCAGGAACA
ATAATCTCACCTCATTACGACTCAATGCTGGTCAAATGCTCATGCTCCGGTTCCACCTAC
GAAATCGTTCGTAGAAAAATGATTCGTGCATTAATCGAGTTCAGAATTAGAGGTGTCAAG
ACCAACATTCCCTTCCTATTGACTCTTTTGACCAATCCAGTATTTATTGAGGGTACATAC
TGGGGGACTTTTATTGACGACACCCCACAACTGTTCCAAATGGTTTCATCACAAAACAGG
GCCCAAAAACTTTTACATTACCTCGCCGACGTGGCAGACAATGGTTCATCTATCAAGGGT
CAAATTGGCTTGCCAAAATTAAAATCAAATCCAAGTGTCCCCCATTTGCACGATGCTCAG
GGCAATGTCATCAACGTTACAAAGTCTGCACCACCATCCGGATGGAGGCAAGTGCTACTA
GAAAAGGGGCCAGCTGAATTTGCCAGACAAGTTAGACAGTTCAATGGTACTTTATTGATG
GACACCACCTGGAGAGACGCTCATCAATCTCTACTTGCAACAAGAGTCAGAACCCACGAT
TTGGCTACAATCGCTCCAACAACCGCACATGCCCTTGCAGGTGCTTTCGCCTTAGAATGT
TGGGGTGGTGCCACATTCGATGTTGCAATGAGATTTTTGCATGAGGATCCATGGCAACGT
TTGAGAAAATTAAGATCTCTGGTGCCTAATATTCCATTCCAAATGTTATTGCGTGGTGCC
AATGGTGTGGCTTATTCTTCATTGCCTGACAATGCTATTGACCATTTCGTCAAGCAAGCC
AAGGATAATAGTGTTGATATATTTAGAGTCTTTGATGCCTTAAATGACTTGGAACAATTG
AAGGTCGGTGTAGATGCTGTGAAGAAGGCAGGTGGTGTTGTAGAAGCCACTGTTTGTTTC
TCTGGGGATATGCTTCAGCCAGGCAAGAAATACAATTTGGATTACTACTTGGAAATTGCT
GAAAAAATTGTCCAAATGGGCACTCATATCCTGGGTATCAAAGATATGGCAGGTACCATG
AAGCCAGCAGCTGCCAAACTACTGATTGGATCTTTGAGGGCTAAGTACCCTGATCTCCCA
ATACATGTTCACACTCACGATTCTGCAGGTACTCGTGTTGCATCAATGACTGCGTGTGCT
CTGGCGGGCGCCGATGTCGTTGATGTTGCCATCAACTCAATGTCTGGTTTAACTTCACAA
CCATCAATCAATGCTCTGTTGGCTTCATTAGAAGGTAATATTGACACTGGTATTAACGTT
GAGCATGTCCGTGAACTAGATGCATATTGGGCAGAGATGAGATTGTTATACTCTTGTTTC
GAGGCTGACTTGAAGGGCCCAGATCCAGAAGTTTATCAACATGAAATCCCAGGTGGTCAA
TTGACAAACTTGTTGTTTCAAGCCCAACAATTGGGTCTTGGAGAACAATGGGCCCAAACA
AAAAGAGCTTACAGAGAAGCCAATTATTTATTGGGTGATATTGTCAAAGTTACCCCAACT
TCGAAGGTCGTTGGTGATCTGGCAAAATTTATGGTCTCCAATAAATTAACTTCCGATGAT
GTGAGACGCCTGGCTAATTCTTTGGATTTCCCTGACTCTGTTATGGATTTCTTCGAAGGC
TTAATCGGCCAACCATATGGTGGGTTCCCAGAACCATTTAGATCAGACGTTTTAAGGAAC
AAGAGAAGAAAGTTGACTTGTCGTCCAGGCCTGGAACTAGAGCCATTTGATCTCGAAAAA
ATTAGAGAAGACTTGCAGAATAGATTTGGTGATGTTGATGAGTGCGACGTTGCTTCTTAT
AACATGTACCCAAGAGTTTATGAAGACTTCCAAAAGATGAGAGAAACGTATGGTGATTTA
TCTGTATTGCCAACAAGAAGCTTTTTGTCTCCACTAGAGACTGACGAAGAAATTGAAGTT
GTAATCGAACAAGGTAAAACGCTAATTATCAAGCTACAGGCTGTGGGTGATTTGAACAAA
AAGACCGGTGAAAGAGAAGTTTACTTTGATTTGAATGGTGAAATGAGAAAAATTCGTGTT
GCTGACAGATCACAAAAAGTGGAAACTGTTACTAAATCCAAAGCAGACATGCATGATCCA
TTACACATTGGTGCACCAATGGCAGGTGTCATTGTTGAAGTTAAAGTTCATAAAGGATCA
CTAATAAAGAAGGGCCAACCTGTAGCCGTATTAAGCGCCATGAAAATGGAAATGATTATA
TCTTCTCCATCCGATGGACAAGTTAAAGAAGTGTTTGTCTCTGATGGTGAAAATGTGGAC
TCTTCTGATTTATTAGTTCTATTAGAAGACCAAGTTCCTGTTGAAACTAAGGCATGA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 1178 |
|---|
| Protein Molecular Weight | 130098.0 |
|---|
| Protein Theoretical pI | 6.15 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >Pyruvate carboxylase 1
MSQRKFAGLRDNFNLLGEKNKILVANRGEIPIRIFRTAHELSMQTVAIYSHEDRLSTHKQ
KADEAYVIGEVGQYTPVGAYLAIDEIISIAQKHQVDFIHPGYGFLSENSEFADKVVKAGI
TWIGPPAEVIDSVGDKVSARNLAAKANVPTVPGTPGPIETVEEALDFVNEYGYPVIIKAA
FGGGGRGMRVVREGDDVADAFQRATSEARTAFGNGTCFVERFLDKPKHIEVQLLADNHGN
VVHLFERDCSVQRRHQKVVEVAPAKTLPREVRDAILTDAVKLAKECGYRNAGTAEFLVDN
QNRHYFIEINPRIQVEHTITEEITGIDIVAAQIQIAAGASLPQLGLFQDKITTRGFAIQC
RITTEDPAKNFQPDTGRIEVYRSAGGNGVRLDGGNAYAGTIISPHYDSMLVKCSCSGSTY
EIVRRKMIRALIEFRIRGVKTNIPFLLTLLTNPVFIEGTYWTTFIDDTPQLFQMVSSQNR
AQKLLHYLADVAVNGSSIKGQIGLPKLKSNPSVPHLHDAQGNVINVTKSAPPSGWRQVLL
EKGPAEFARQVRQFNGTLLMDTTWRDAHQSLLATRVRTHDLATIAPTTAHALAGRFALEC
WGGATFDVAMRFLHEDPWERLRKLRSLVPNIPFQMLLRGANGVAYSSLPDNAIDHFVKQA
KDNGVDIFRVFDALNDLEQLKVGVDAVKKAGGVVEATVCFSGDMLQPGKKYNLDYYLEIA
EKIVQMGTHILGIKDMAGTMKPAAAKLLIGSLRAKYPDLPIHVHTHDSAGTAVASMTACA
LAGADVVDVAINSMSGLTSQPSINALLASLEGNIDTGINVEHVRELDAYWAEMRLLYSCF
EADLKGPDPEVYQHEIPGGQLTNLLFQAQQLGLGEQWAETKRAYREANYLLGDIVKVTPT
SKVVGDLAQFMVSNKLTSDDVRRLANSLDFPDSVMDFFEGLIGQPYGGFPEPFRSDVLRN
KRRKLTCRPGLELEPFDLEKIREDLQNRFGDVDECDVASYNMYPRVYEDFQKMRETYGDL
SVLPTRSFLSPLETDEEIEVVIEQGKTLIIKLQAVGDLNKKTGEREVYFDLNGEMRKIRV
ADRSQKVETVTKSKADMHDPLHIGAPMAGVIVEVKVHKGSLIKKGQPVAVLSAMKMEMII
SSPSDGQVKEVFVSDGENVDSSDLLVLLEDQVPVETKA |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Lim, F., Morris, C. P., Occhiodoro, F., Wallace, J. C. (1988). "Sequence and domain structure of yeast pyruvate carboxylase." J Biol Chem 263:11493-11497.3042770
- Feuermann, M., de Montigny, J., Potier, S., Souciet, J. L. (1997). "The characterization of two new clusters of duplicated genes suggests a 'Lego' organization of the yeast Saccharomyces cerevisiae chromosomes." Yeast 13:861-869.9234674
- Tettelin, H., Agostoni Carbone, M. L., Albermann, K., Albers, M., Arroyo, J., Backes, U., Barreiros, T., Bertani, I., Bjourson, A. J., Bruckner, M., Bruschi, C. V., Carignani, G., Castagnoli, L., Cerdan, E., Clemente, M. L., Coblenz, A., Coglievina, M., Coissac, E., Defoor, E., Del Bino, S., Delius, H., Delneri, D., de Wergifosse, P., Dujon, B., Kleine, K., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome VII." Nature 387:81-84.9169869
- Morris, C. P., Lim, F., Wallace, J. C. (1987). "Yeast pyruvate carboxylase: gene isolation." Biochem Biophys Res Commun 145:390-396.3036126
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
|
|---|