| Identification |
|---|
| Name | CDP-diacylglycerol--inositol 3-phosphatidyltransferase |
|---|
| Synonyms | - Phosphatidylinositol synthase
- PI synthase
- PtdIns synthase
|
|---|
| Gene Name | PIS1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in phosphotransferase activity, for other substituted phosphate groups |
|---|
| Specific Function | CDP-diacylglycerol + myo-inositol = CMP + phosphatidyl-1D-myo-inositol |
|---|
| Cellular Location | Microsome membrane; Multi-pass membrane protein. Endoplasmic reticulum membrane; Multi-pass membrane protein. Mitochondrion outer membrane |
|---|
| SMPDB Pathways | |
|---|
| KEGG Pathways | | Glycerophospholipid metabolism | ec00564 | 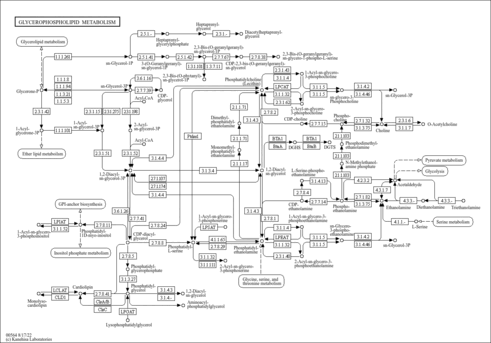 | | Inositol phosphate metabolism | ec00562 | 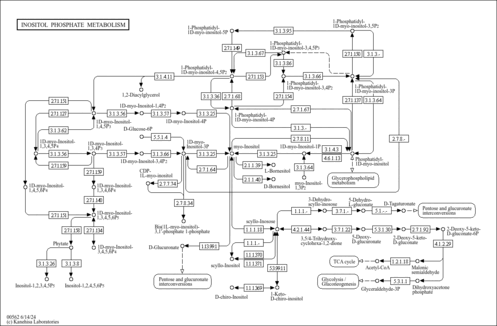 |
|
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00173 | Myoinositol | Show | | YMDB00230 | 1-Phosphatidyl-D-myo-inositol | Show | | YMDB00406 | Cytidine monophosphate | Show | | YMDB00862 | hydron | Show |
|
|---|
| GO Classification | | Component |
|---|
| cell part | | membrane | | Function |
|---|
| transferase activity, transferring phosphorus-containing groups | | phosphotransferase activity, for other substituted phosphate groups | | catalytic activity | | transferase activity | | Process |
|---|
| organophosphate metabolic process | | phospholipid metabolic process | | phospholipid biosynthetic process | | metabolic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 16 |
|---|
| Locus | YPR113W |
|---|
| Gene Sequence | >663 bp
ATGAGTTCGAATTCAACACCAGAAAAGGTTACTGCAGAACACGTTCTGTGGTATATTCCC
AATAAGATCGGTTATGTTCGTGTTATCACCGCCGCCCTTTCTTTTTTCGTTATGAAGAAT
CATCCTACGGCCTTTACATGGTTGTATAGTACATCATGTCTACTGGATGCGCTAGACGGA
ACCATGGCAAGAAAGTACAATCAGGTTTCCAGTCTGGGTGCCGTTCTGGACATGGTTACC
GACAGATCCAGTACCGCTGGCTTGATGTGTTTCCTTTGTGTGCAGTATCCCCAATGGTGC
GTTTTCTTCCAATTAATGCTGGGCTTGGATATTACTAGTCACTACATGCATATGTATGCC
AGTTTAAGTGCTGGTAAGACTTCTCATAAAAGTGTGGGCGAGGGTGAGTCCAGATTGTTA
CACCTGTACTACACGAGAAGAGACGTACTGTTCACTATCTGTGCGTTTAACGAACTATTT
TATGCTGGATTGTACTTGCAGTTGTTCTCAAATTCTGCAACCTTTGGTAAATGGACTACA
ATCATAAGTTTCCCTGGTTACGTGTTCAAGCAGACCGCAAACGTTGTCCAGCTAAAAAGG
GCAGCCTTGATTTTAGCAGACAACGATGCCAAGAATGCCAACGAGAAGAACAAGACTTAC
TGA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 220 |
|---|
| Protein Molecular Weight | 24823.5 |
|---|
| Protein Theoretical pI | 8.87 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | - 21-41
- 46-66
- 76-96
- 99-119
- 146-166
- 171-191
|
|---|
| Protein Sequence | >CDP-diacylglycerol--inositol 3-phosphatidyltransferase
MSSNSTPEKVTAEHVLWYIPNKIGYVRVITAALSFFVMKNHPTAFTWLYSTSCLLDALDG
TMARKYNQVSSLGAVLDMVTDRSSTAGLMCFLCVQYPQWCVFFQLMLGLDITSHYMHMYA
SLSAGKTSHKSVGEGESRLLHLYYTRRDVLFTICAFNELFYAGLYLQLFSNSATFGKWTT
IISFPGYVFKQTANVVQLKRAALILADNDAKNANEKNKTY |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Nikawa, J., Kodaki, T., Yamashita, S. (1987). "Primary structure and disruption of the phosphatidylinositol synthase gene of Saccharomyces cerevisiae." J Biol Chem 262:4876-4881.3031032
- Bussey, H., Storms, R. K., Ahmed, A., Albermann, K., Allen, E., Ansorge, W., Araujo, R., Aparicio, A., Barrell, B., Badcock, K., Benes, V., Botstein, D., Bowman, S., Bruckner, M., Carpenter, J., Cherry, J. M., Chung, E., Churcher, C., Coster, F., Davis, K., Davis, R. W., Dietrich, F. S., Delius, H., DiPaolo, T., Hani, J., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome XVI." Nature 387:103-105.9169875
- Ghaemmaghami, S., Huh, W. K., Bower, K., Howson, R. W., Belle, A., Dephoure, N., O'Shea, E. K., Weissman, J. S. (2003). "Global analysis of protein expression in yeast." Nature 425:737-741.14562106
- Kim, H., Melen, K., Osterberg, M., von Heijne, G. (2006). "A global topology map of the Saccharomyces cerevisiae membrane proteome." Proc Natl Acad Sci U S A 103:11142-11147.16847258
|
|---|

