| Identification |
|---|
| Name | ATP phosphoribosyltransferase |
|---|
| Synonyms | |
|---|
| Gene Name | HIS1 |
|---|
| Enzyme Class | |
|---|
| Biological Properties |
|---|
| General Function | Involved in ATP phosphoribosyltransferase activity |
|---|
| Specific Function | Catalyzes the condensation of ATP and 5-phosphoribose 1- diphosphate to form N'-(5'-phosphoribosyl)-ATP (PR-ATP). Has a crucial role in the pathway because the rate of histidine biosynthesis seems to be controlled primarily by regulation of the enzymatic activity |
|---|
| Cellular Location | Cytoplasm |
|---|
| SMPDB Pathways | |
|---|
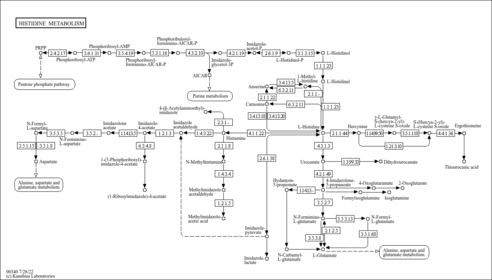
| KEGG Pathways | |
|---|
| SMPDB Reactions | |
|---|
| KEGG Reactions | |
|---|
| Metabolites | | YMDB ID | Name | View |
|---|
| YMDB00109 | Adenosine triphosphate | Show | | YMDB00165 | Phosphoribosyl-ATP | Show | | YMDB00219 | Pyrophosphate | Show | | YMDB00402 | Phosphoribosyl pyrophosphate | Show | | YMDB00862 | hydron | Show | | YMDB00868 | 1-(5-phosphoribosyl)-5'-AMP | Show | | YMDB16274 | 1-(5-phosphoribosyl)-ATP | Show |
|
|---|
| GO Classification | | Component |
|---|
| cell part | | intracellular part | | cytoplasm | | Function |
|---|
| ion binding | | catalytic activity | | cation binding | | transferase activity | | metal ion binding | | transferase activity, transferring glycosyl groups | | magnesium ion binding | | ATP phosphoribosyltransferase activity | | binding | | transferase activity, transferring pentosyl groups | | Process |
|---|
| metabolic process | | cellular metabolic process | | cellular amino acid and derivative metabolic process | | cellular amino acid metabolic process | | histidine family amino acid metabolic process | | histidine metabolic process | | histidine biosynthetic process |
|
|---|
| Gene Properties |
|---|
| Chromosome Location | chromosome 5 |
|---|
| Locus | YER055C |
|---|
| Gene Sequence | >894 bp
ATGGATTTGGTGAACCATCTAACCGATAGACTACTGTTTGCAATCCCAAAGAAAGGTCGT
TTATATTCTAAAAGTGTTTCTATTTTGAATGGTGCTGATATTACCTTTCACCGCTCTCAA
AGATTAGACATTGCACTAAGCACAAGCTTACCTGTAGCGTTGGTCTTTCTGCCCGCTGCA
GATATTCCAACTTTTGTTGGTGAAGGTAAATGTGATCTTGGTATAACTGGTGTTGACCAA
GTTCGTGAATCTAACGTCGACGTAGACTTAGCAATCGATTTGCAATTTGGTAACTGTAAA
TTGCAGGTACAAGTCCCCGTAAATGGCGAGTATAAAAAGCCAGAACAGTTAATTGGCAAA
ACCATTGTTACCAGTTTCGTGAAACTTGCTGAAAAATACTTTGCCGATTTGGAAGGTACT
ACTGTTGAAAAAATGACCACAAGGATAAAGTTTGTCAGTGGTTCCGTGGAGGCATCATGT
GCTCTGGGAATTGGTGATGCTATTGTAGATCTTGTAGAGAGTGGTGAGACAATGAGGGCA
GCAGGTTTAGTTGATATTGCCACCGTCCTAAGCACAAGTGCCTACCTAATAGAATCAAAG
AACCCAAAGAGCGATAAGAGTTTGATTGCTACTATCAAATCAAGAATTGAAGGTGTCATG
ACCGCTCAAAGGTTCGTTTCATGTATTTATAACGCACCTGAAGACAAGCTGCCTGAACTG
TTGAAGGTGACGCCTGGCCGTAGAGCACCAACCATTTCCAAAATTGACGATGAAGGATGG
GTTGCTGTTAGTTCCATGATTGAGAGAAAAACGAAGGGTGTTGTTTTAGATGAATTGAAA
AGACTCGGCGCATCTGATATCATGGTTTTCGAAATTTCTAATTGTCGTGTATAA |
|---|
| Protein Properties |
|---|
| Pfam Domain Function | |
|---|
| Protein Residues | 297 |
|---|
| Protein Molecular Weight | 32266.09961 |
|---|
| Protein Theoretical pI | 5.94 |
|---|
| Signalling Regions | |
|---|
| Transmembrane Regions | |
|---|
| Protein Sequence | >ATP phosphoribosyltransferase
MDLVNHLTDRLLFAIPKKGRLYSKSVSILNGADITFHRSQRLDIALSTSLPVALVFLPAA
DIPTFVGEGKCDLGITGVDQVRESNVDVDLAIDLQFGNCKLQVQVPVNGEYKKPEQLIGK
TIVTSFVKLAEKYFADLEGTTVEKMTTRIKFVSGSVEASCALGIGDAIVDLVESGETMRA
AGLVDIATVLSTSAYLIESKNPKSDKSLIATIKSRIEGVMTAQRFVSCIYNAPEDKLPEL
LKVTPGRRAPTISKIDDEGWVAVSSMIERKTKGVVLDELKRLGASDIMVFEISNCRV |
|---|
| References |
|---|
| External Links | |
|---|
| General Reference | - Hinnebusch, A. G., Fink, G. R. (1983). "Repeated DNA sequences upstream from HIS1 also occur at several other co-regulated genes in Saccharomyces cerevisiae." J Biol Chem 258:5238-5247.6300123
- Dietrich, F. S., Mulligan, J., Hennessy, K., Yelton, M. A., Allen, E., Araujo, R., Aviles, E., Berno, A., Brennan, T., Carpenter, J., Chen, E., Cherry, J. M., Chung, E., Duncan, M., Guzman, E., Hartzell, G., Hunicke-Smith, S., Hyman, R. W., Kayser, A., Komp, C., Lashkari, D., Lew, H., Lin, D., Mosedale, D., Davis, R. W., et, a. l. .. (1997). "The nucleotide sequence of Saccharomyces cerevisiae chromosome V." Nature 387:78-81.9169868
|
|---|